Rank-Based Regression in R: Rfit Package for Robust Linear Models
Rank-based regression fits a linear model by minimising a function of the ranks of the residuals instead of their squared values. R's Rfit package implements this with a near drop-in replacement for lm() that resists outliers in the response and stays efficient when errors are non-Normal.
install.packages("Rfit") once in R, then the code blocks below will work as shown. The #> comments display the output you should see.When does a single outlier ruin a regression?
Ordinary least squares squares every residual before adding them up, so one giant residual from a single bad point can dominate the loss and drag the fitted line toward that point. Rank-based regression replaces the squared-residual loss with a function of the residual ranks, so the worst residual contributes at most rank n, not n². The cleanest way to feel that difference is to take a clean line, knock one y-value far off, and fit both methods side by side.
The true slope is 2 and the true intercept is 0. OLS reports an intercept of 4.93, pulled almost five units upward by that one contaminated observation. The rank-based fit reports 0.31, a hair off zero, because the bad point's residual gets capped at rank 30 instead of being squared. The slope barely budges either way; intercepts move first because they absorb shifts in the response.
Try it: Add a second bad point at index 25 (y_dirty[25] <- y_dirty[25] - 60) and refit both. Does OLS lose its slope this time? Does rfit()?
Click to reveal solution
Explanation: Two outliers in opposite directions don't cancel under OLS, they amplify the slope error. The rank loss bounds each contribution, so two bad points are no worse than one.
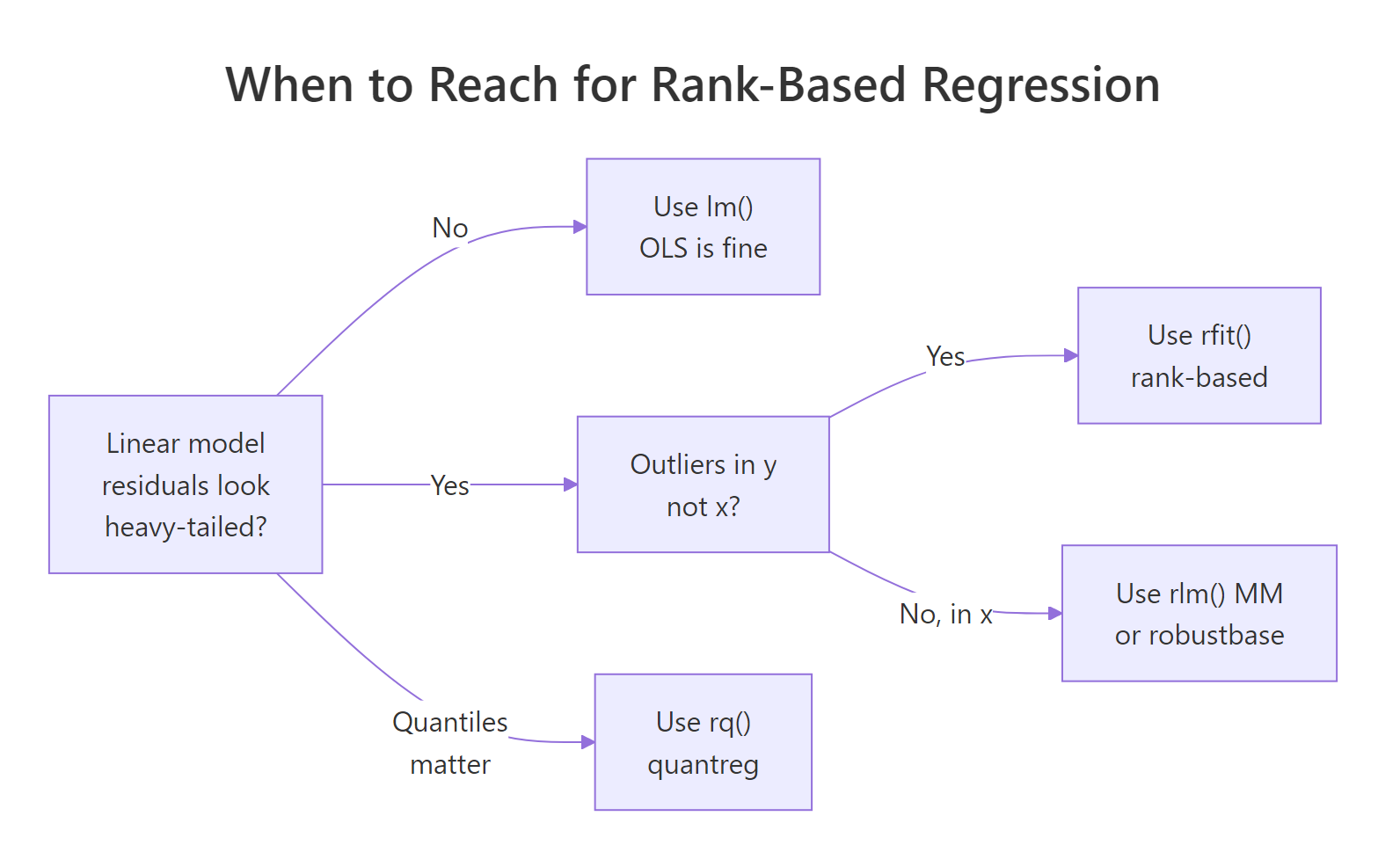
Figure 1: When rank-based regression is the right tool, and when a different robust method fits better.
How does rfit() syntax compare to lm()?
rfit() takes the same formula interface as lm(). Anything you can write on the left of ~ and the right of ~ for lm(), you can hand to rfit() unchanged: predictors, interactions, factor variables, polynomial terms. The only thing that changes is the loss function under the hood. That makes the typical workflow "swap lm for rfit and re-read the summary."
Three things to notice. First, the coefficient table looks just like lm's, with rank-based standard errors instead of OLS ones. Second, the Multiple R-squared (Robust) is the rank-based analogue of R²; treat it the same way you'd treat OLS R² (higher is more variance explained). Third, the Reduction in Dispersion Test is the rank-based F-test for "is at least one slope nonzero?" The 48.06 statistic with effectively zero p-value says yes, weight and horsepower together explain mpg.
lm summary's F-statistic to flag a useful model, you can trust the dispersion test the same way for an rfit summary.Try it: Fit rfit() on airquality predicting Ozone from Temp and Wind. Drop missing rows first with na.omit(). Read the dispersion test p-value.
Click to reveal solution
Explanation: Hot still days drive ozone up. Both effects are highly significant under rank-based inference, and the robust R² of about 0.50 matches what an OLS fit on this data would also give, since airquality is fairly well behaved.
What score function should you choose?
The "rank-based" loss is more precisely a score-weighted sum of ranked residuals. Different score functions give different trade-offs between efficiency on Normal data and robustness to heavy tails. Rfit ships several; you pick one with the scores = argument.
The default is Wilcoxon scores (wscores). They are optimal when errors are logistic-distributed (heavy-ish tails) and only 4.5% less efficient than OLS when errors are actually Normal. That 95.5% asymptotic relative efficiency is what makes rank-based regression a near-free upgrade for general use.
The Jaeckel dispersion that Wilcoxon scores minimise is
$$D(\boldsymbol{\beta}) = \sum_{i=1}^{n} \varphi\!\left(\frac{R(e_i)}{n+1}\right) e_i$$
Where:
- $e_i = y_i - \mathbf{x}_i^\top \boldsymbol{\beta}$ is the i-th residual
- $R(e_i)$ is the rank of $e_i$ among all residuals
- $\varphi(u) = \sqrt{12}\,(u - \tfrac{1}{2})$ is the Wilcoxon score function
For very heavy tails (think Cauchy-like), sign scores (sgnscores) bound each contribution even harder. For right-skewed errors, bent scores are designed to handle the asymmetry. The diagram below maps the choice.
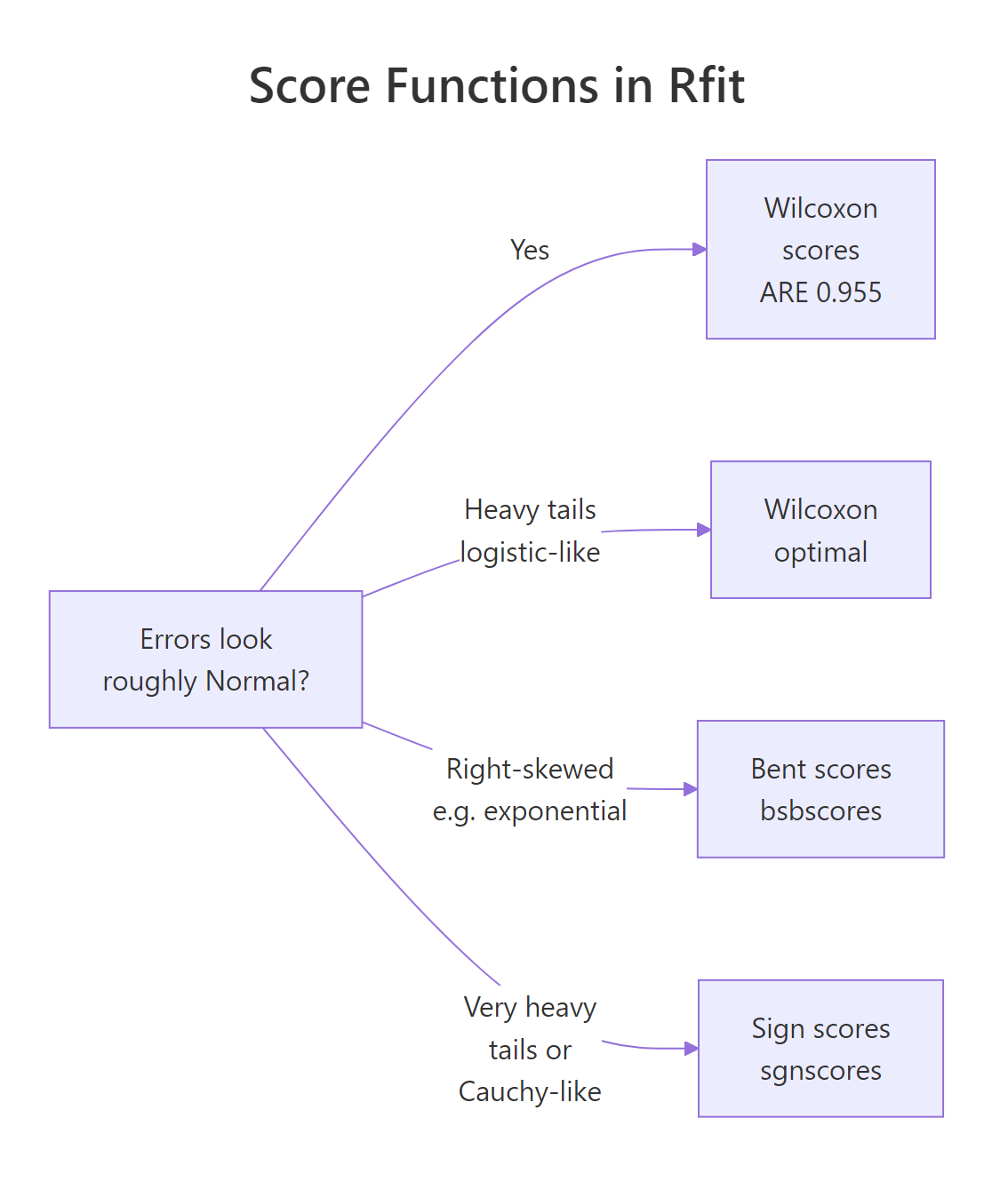
Figure 2: Pick a score function from what you know about the error distribution, not from the data.
Both estimators recover the true slope of 2, but their standard errors differ. Sign scores have a slight efficiency edge here because the t with 2 df has very heavy tails (its theoretical variance is infinite). Wilcoxon still does respectably; it is rarely the worst choice across error distributions, which is why it's the default.
Try it: Refit the original contaminated y_dirty ~ x line with scores = sgnscores and compare its slope to the Wilcoxon fit from earlier.
Click to reveal solution
Explanation: With one severe outlier, sign and Wilcoxon scores give essentially the same slope, because both bound the bad point's contribution. The choice matters more when the whole error distribution is heavy-tailed, not when one point misbehaves.
How do you test nested models with drop.test()?
To compare two nested rank-based models you use drop.test(full, reduced). It's the rank-based answer to anova(lm1, lm2). The test statistic is a reduction-in-dispersion F, and its p-value tells you whether the dropped predictors mattered.
The dropped term is cyl. The reduction-in-dispersion F is 2.75 with p = 0.11. At a 5% threshold you can't reject the simpler model. Adding cyl on top of wt and hp doesn't significantly tighten the rank-based fit. That matches the OLS conclusion on the same data, but the rank-based test would still hold up if a few mpg values had been corrupted.
drop.test() works exactly like anova() for nested OLS models. Pass the full model first, the reduced model second. If you swap the order, the F-statistic flips sign in derivation but the p-value is unchanged.Try it: Build a nested pair where the reduced model drops wt instead of cyl. Does dropping weight cost more than dropping cylinders?
Click to reveal solution
Explanation: Weight carries a huge share of the explanation in mtcars. Removing it triples the dispersion, and the rank-based F flags that with a p-value far below 0.001. That asymmetry, wt critical and cyl borderline, is exactly the kind of conclusion you want a robust test to reach reliably.
How do you run ANOVA with Rfit?
For categorical predictors, Rfit gives you oneway.rfit() for one-way designs and raov() for multi-way / factorial layouts. Both wrap the same dispersion machinery as rfit() and produce a familiar ANOVA table.
The test statistic and p-value answer one question: are the three species means (well, locations, since rank-based methods speak in shift, not means) different? The 91.3 statistic with p ≈ 0 says yes, decisively. It's the rank-based parallel of aov(Sepal.Length ~ Species, data = iris), but resistant to a few stray flowers with mismeasured petals.
For factorial designs, raov() accepts a formula:
Both factors have small p-values; dose carries more of the rank-based explanatory weight (larger reduction in dispersion per df). Read the table the way you'd read a base anova() table, swapping "Mean Sq" for "Mean RD" (mean reduction in dispersion).
Try it: Run raov(weight ~ group, data = PlantGrowth). Is group significant?
Click to reveal solution
Explanation: The treatments differ at the 5% level under rank-based inference, mirroring the classical aov() conclusion on this dataset.
How do you check a rank-based fit?
A robust fit doesn't excuse you from diagnostics. You still want to spot leverage, structural misspecification, and any residual pattern. Rfit provides studentised residuals through a method dispatch on rstudent(), and you plot them against fitted values just as you would for lm.
One observation pokes a little above +2 (the Toyota Corolla, a famously light, fuel-efficient car the model can't fully predict). Nothing is dramatically beyond the ±2 guidelines, so the rank-based fit isn't hiding leverage drama, just a few honest tail points. If you saw a clear funnel or curve in this plot, that would tell you to revisit the model form, not just to switch to a robust loss.
rfit() only protects against outliers in y, not against fitting the wrong shape.Try it: Find the row index of the largest absolute studentised residual.
Click to reveal solution
Explanation: The Corolla is an over-performer; its actual mpg sits well above what wt + hp + cyl predicts. A rank-based fit doesn't try to chase it the way OLS would, but the diagnostic still flags it for follow-up.
Practice Exercises
These combine pieces from across the tutorial. Use distinct variable names so the inline state stays clean.
Exercise 1: Contaminate airquality and compare slopes
Load airquality, drop missing values, then push Ozone[1] to 500 (a deliberate outlier). Fit both lm() and rfit() for Ozone ~ Temp + Wind and report the difference in the Temp slope.
Click to reveal solution
Explanation: One contaminated point has dragged the OLS Temp slope from about 1.62 down to 1.35, a ~17% bias. The rank-based slope is right where the clean fit said it should be. This is the practical payoff of swapping lm for rfit on data you don't fully trust.
Exercise 2: Score choice on Cauchy errors
Simulate n = 200 observations from y = 3 + 0.5 * x + cauchy_noise where x runs uniform on [0, 20]. Fit rfit() twice (Wilcoxon and sign scores) and report which gives a smaller standard error on the slope.
Click to reveal solution
Explanation: Sign scores cut the slope's standard error roughly in half on Cauchy-tailed data, because they bound every residual contribution at ±1 instead of letting moderate residuals contribute proportionally to their rank. When tails are this heavy, the sign loss is dramatically more efficient than Wilcoxon, and worlds better than OLS, which has no defined variance to estimate at all.
Complete Example
Putting all six pieces together on mtcars: contaminate, fit, test, diagnose.
Despite the planted Mazda outlier, the slopes match the clean-data conclusions from the earlier sections almost exactly. drop.test() still says cyl is borderline (p = 0.11). The diagnostic correctly flags the Mazda as the worst residual without letting it deform the fit. An OLS pass on the same mt2 would have produced visibly different slopes and possibly flipped the cyl significance call.
Summary
| Task | Function | Notes |
|---|---|---|
| Fit a robust linear model | rfit(y ~ x, data = df) |
lm-style formula |
| Read coefficients & robust R² | summary(fit) |
Familiar layout |
| Compare nested models | drop.test(full, reduced) |
Rank-based F-test |
| One-way ANOVA | oneway.rfit(y, group) |
Resistant to outliers |
| Multi-way / factorial ANOVA | raov(y ~ a + b, data = df) |
"Mean RD" instead of "Mean Sq" |
| Diagnose | rstudent(fit), fitted(fit) |
Plot residuals vs fits |
| Switch loss function | scores = wscores / sgnscores / bbscores |
Wilcoxon by default |
Key takeaways:
- Rank-based regression bounds each residual's influence by its rank, not its square, so a single bad y-value can't capture the line.
- Wilcoxon scores cost you ~4.5% efficiency on clean Normal data and pay you back many times over on heavier-tailed or contaminated data.
- The summary, dispersion test, and ANOVA tables all read like their classical counterparts, so adoption is mostly a matter of swapping
lmforrfit. - Robust loss does not fix structural misspecification. You still need diagnostics.
References
- Kloke, J. D. & McKean, J. W. (2012). Rfit: Rank-based Estimation for Linear Models. The R Journal, 4(2), 57–64. Link
- Hettmansperger, T. P. & McKean, J. W. (2011). Robust Nonparametric Statistical Methods, 2nd ed. CRC Press.
- Jaeckel, L. A. (1972). Estimating regression coefficients by minimizing the dispersion of the residuals. Annals of Mathematical Statistics, 43, 1449–1458.
- Rfit on CRAN. Link
- Rfit reference manual. Link
- Mangiafico, S. (2016). R Companion: Nonparametric Regression. Link
- Kloke, J. Rfit GitHub repository. Link
Continue Learning
- Quantile Regression in R: quantreg Package. Model any conditional quantile, not just the mean. Sister method to rank-based regression for non-Normal y.
- Robust Regression in R. M-estimators via
MASS::rlm, the other major robust toolkit. Use when leverage points (bad x) are the worry, not just bad y. - Wilcoxon, Mann-Whitney and Kruskal-Wallis in R. Two-sample and k-sample rank tests, the conceptual ancestors of the loss
rfit()minimises.