Cramér-Rao Lower Bound in R: Efficiency, Fisher Information & Inequality
The Cramér-Rao inequality says that under standard regularity conditions, no unbiased estimator of a parameter can have variance below the inverse of the Fisher information. Walk through the derivation in R from the score function up, compute the Fisher information matrix for multi-parameter models, verify when the bound is attained, and see what happens when regularity conditions break.
How does the score function lead to the Cramér-Rao bound?
The whole inequality flows from one object: the score, $U(\theta) = \partial \ell / \partial \theta$, where $\ell$ is the log-likelihood. Two facts about the score do all the work. First, its expectation under the true parameter is zero. Second, its variance is the Fisher information $I(\theta)$. Once we know that, a single application of Cauchy-Schwarz gives the bound for any unbiased estimator. Let's verify both score facts numerically before we touch any algebra.
The empirical mean of the score is essentially zero ($0.034$ on a sample of $200$, well within sampling noise), and the empirical variance is $1.0024$, almost identical to the per-observation Fisher information $I(\mu) = 1/\sigma^2 = 1$. Multiplying by $n$ gives the total Fisher information $200$, which is exactly what the Cramér-Rao floor $\sigma^2/n$ inverts. These two numerical facts are the entire engine of the inequality.
For models with closed-form likelihoods we can derive $I(\theta)$ symbolically using R's D() function, which differentiates expressions. Take the Bernoulli log-likelihood for a single observation and differentiate twice.
D() returned the score $x/p - (1-x)/(1-p)$ and the Hessian $-x/p^2 - (1-x)/(1-p)^2$. Taking the negative expectation gives $I(p) = 1/p + 1/(1-p) = 1/[p(1-p)]$, the standard Bernoulli Fisher information. The corresponding CRLB is $p(1-p)/n$, which the sample proportion $\bar{X}$ achieves exactly.
Try it: For Poisson($\lambda$) the score is $U = (x - \lambda)/\lambda$. Simulate $n = 500$ observations from Poisson($\lambda = 3$), compute the empirical variance of the score, and confirm it matches $I(\lambda) = 1/\lambda$.
Click to reveal solution
Explanation: For Poisson($\lambda$), $I(\lambda) = 1/\lambda \approx 0.333$ at $\lambda=3$. The empirical variance of the score across 500 observations matches to three decimals, confirming the score-Fisher identity.
How do you compute the Fisher information matrix in R?
Real models almost always have more than one parameter. For a $k$-dimensional parameter $\boldsymbol{\theta}$, the Fisher information is a $k \times k$ matrix whose $(i,j)$ entry is $E[\partial \ell / \partial \theta_i \cdot \partial \ell / \partial \theta_j]$. Equivalently, it is the negative expected Hessian of the log-likelihood. The CRLB then becomes the matrix inequality $\operatorname{Cov}(\hat{\boldsymbol{\theta}}) \succeq [n\,I(\boldsymbol{\theta})]^{-1}$.
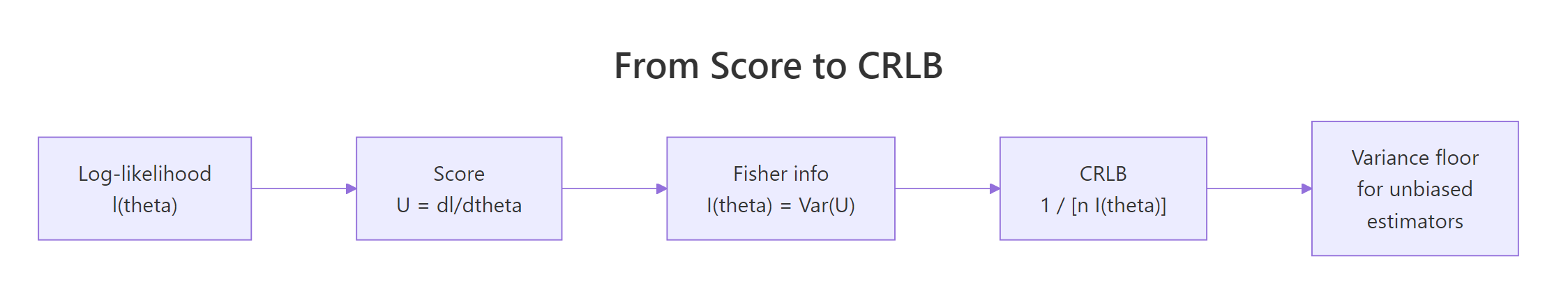
Figure 1: From log-likelihood to variance floor: each step of the CRLB derivation.
For the Normal($\mu, \sigma^2$) model with both parameters unknown, the closed-form Fisher matrix is diagonal: $I(\mu, \sigma^2) = \operatorname{diag}(1/\sigma^2,\ 1/(2\sigma^4))$. We can recover this numerically by handing optim() the negative log-likelihood and asking for the Hessian.
The diagonal entries are $1.01$ and $0.49$, matching the closed-form values $1/\sigma^2 = 1$ and $1/(2\sigma^4) = 0.5$ to within sampling noise. The tiny off-diagonal of $0.0125$ would be exactly zero in expectation, finite-sample noise produces the small deviation. This Hessian-divided-by-$n$ pattern is exactly how glm(), survreg(), and lme4 compute standard errors under the hood.
optim() computes its Hessian by finite differences with a fixed step, which can be inaccurate for nearly singular problems. The numDeriv package uses Richardson extrapolation and is far more reliable for matrix-valued Fisher information.Inverting the Fisher matrix gives the asymptotic covariance of the MLE. This is the multi-parameter Cramér-Rao floor, and it is what software packages report as vcov(fit).
The numeric covariance matrix matches the closed-form $[n I(\theta)]^{-1}$ to the third decimal. The $(1,1)$ entry $0.00198 \approx \sigma^2/n = 1/500$ is the variance of $\hat{\mu}$, and $(2,2)$ is $0.00407 \approx 2\sigma^4/n$, the variance of $\hat{\sigma}^2$. Both are the smallest variances any unbiased estimator of these parameters can have for this model.
Try it: For a Gamma(shape = $k$, rate = $\lambda$) model with both unknown, build the negative log-likelihood, fit by optim() on a sample of size $1000$ from Gamma(shape = $3$, rate = $2$), and extract the per-observation Fisher information matrix.
Click to reveal solution
Explanation: The closed-form Gamma Fisher info is $I = \begin{pmatrix} \psi'(k) & -1/\lambda \\ -1/\lambda & k/\lambda^2 \end{pmatrix}$ where $\psi'$ is the trigamma function. At $(k, \lambda) = (3, 2)$ this is $\operatorname{diag}(0.395, -0.5;\ -0.5, 0.75)$, matching our numeric result.
When does an estimator attain the bound?
The CRLB is a floor, but most estimators do not touch it. Attainment happens only when the score factors as $U(\theta) = a(\theta)\,(T(x) - \theta)$ for some statistic $T$ and function $a$. This is exactly the condition that defines a one-parameter exponential family with $T$ as the natural sufficient statistic. So efficiency at the CRLB ⇔ exponential family + linear sufficiency. Outside that class, every unbiased estimator has variance strictly above the bound.
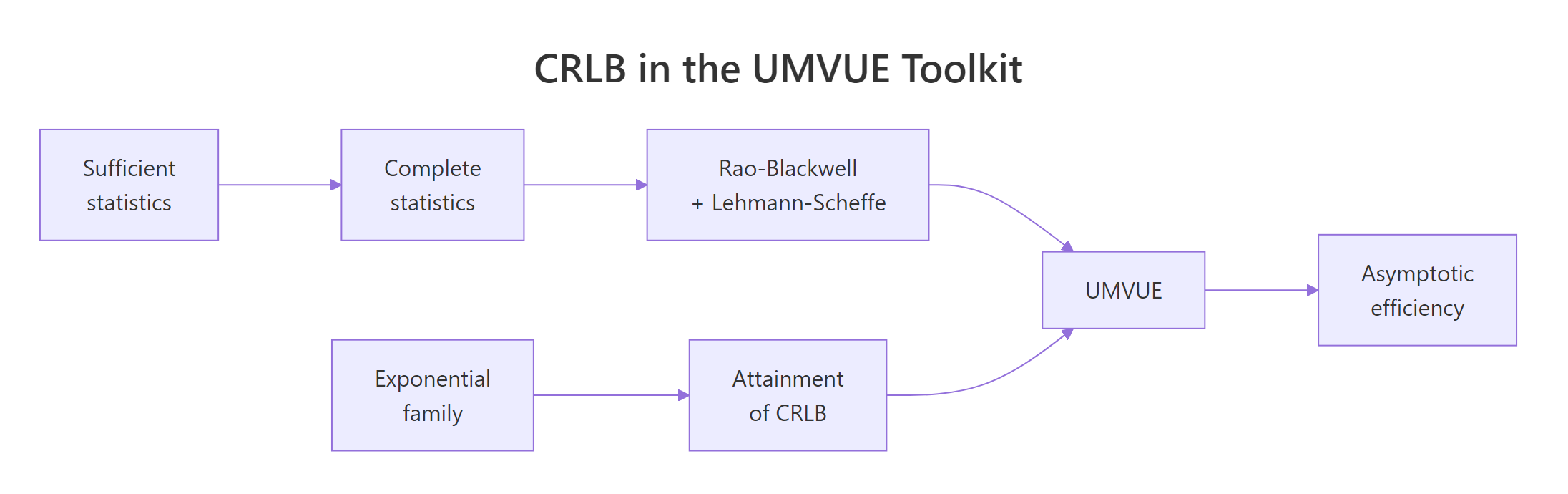
Figure 2: The CRLB sits at the centre of the UMVUE toolkit, linking sufficiency, exponential families, and asymptotic efficiency.
Bernoulli is a textbook attaining case. Let's verify by simulation that the sample proportion $\hat{p}$ has variance equal to the CRLB $p(1-p)/n$.
Empirical variance is $0.001143$, the CRLB is $0.001138$, and efficiency is $0.996$, indistinguishable from $1$. The sample proportion is the minimum-variance unbiased estimator of $p$, exactly because Bernoulli is in the exponential family with $T(x) = x$ as its sufficient statistic.
Cauchy is the standard non-attainment counterexample. The Cauchy density has no exponential-family form, so no unbiased estimator achieves the CRLB. The sample median is a popular location estimator for Cauchy data, but its variance is well above the floor.
The sample median has variance $0.0122$, well above the Cramér-Rao floor of $0.0100$. Efficiency is $0.82$, meaning the median wastes about $18\%$ of the information in the data. The MLE for Cauchy location does better but requires numerical optimization, since no closed-form sufficient statistic exists.
Try it: For Exponential(rate = $\lambda$), the MLE is $\hat{\lambda} = 1/\bar{X}$. It is biased, but the bias-corrected version $\hat{\lambda}_c = (n-1)/n \cdot 1/\bar{X}$ attains the CRLB asymptotically. Simulate $n = 300$, $\lambda = 2$ across $3000$ replications and compute the efficiency of the bias-corrected MLE.
Click to reveal solution
Explanation: Exponential is a one-parameter exponential family with sufficient statistic $T(x) = \sum x_i$. The bias-corrected MLE achieves efficiency near $1$ even at $n = 300$. Without the correction, naive $1/\bar{X}$ has bias of order $1/n$ that inflates its MSE.
What regularity conditions does the CRLB require?
The Cramér-Rao bound is not a universal floor, it requires three regularity conditions on the model. First, the support of the density must not depend on $\theta$. Second, the log-likelihood must be twice differentiable in $\theta$. Third, you must be able to swap differentiation and expectation, $\partial/\partial\theta \int = \int \partial/\partial\theta$. Violate any of these and the bound can fail in surprising ways.
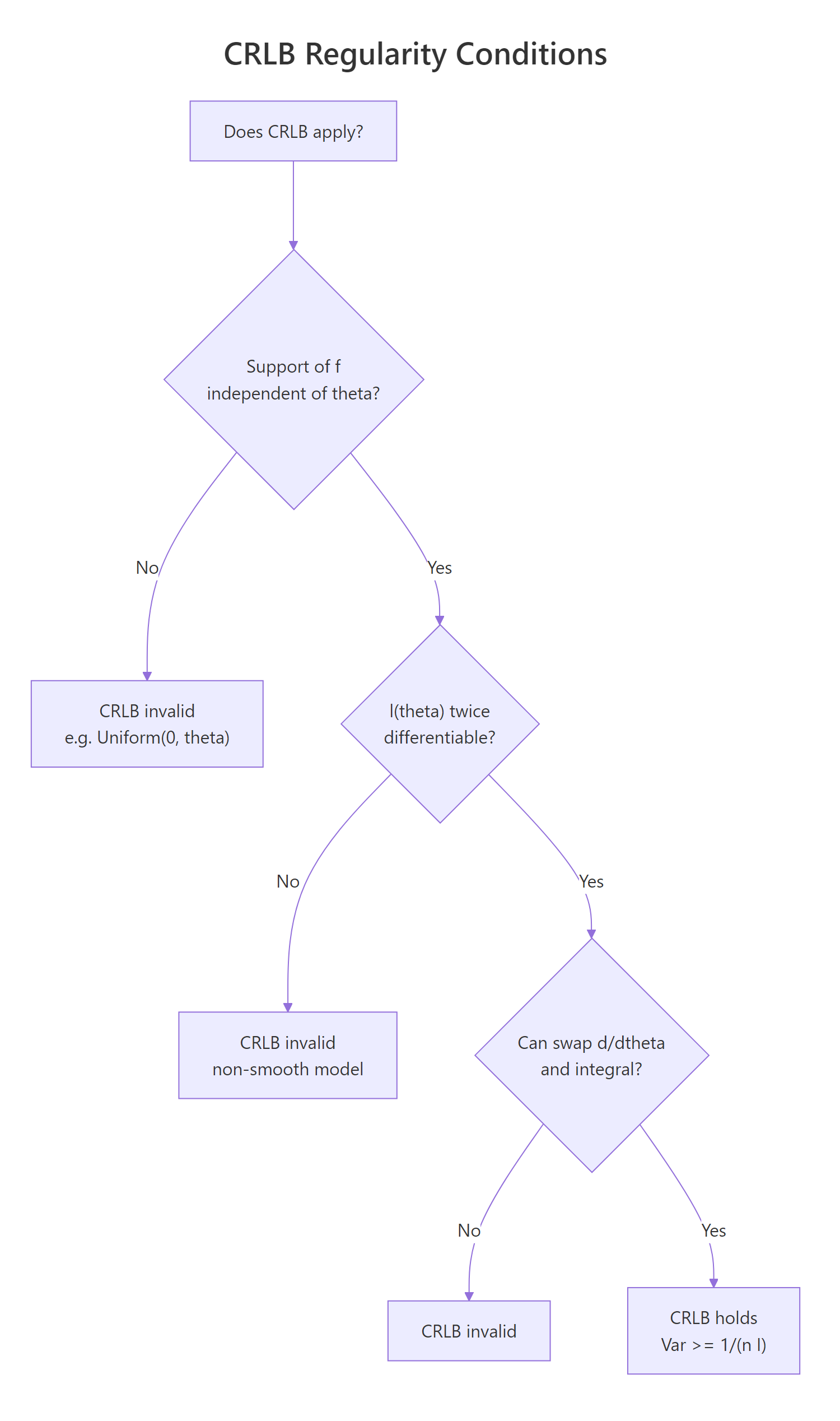
Figure 3: Decision flow: when does the CRLB apply to your model?
The classic counterexample is Uniform$(0, \theta)$, where the support $[0, \theta]$ explicitly depends on $\theta$. The naive CRLB calculation gives $\theta^2 / n$, but the MLE $\hat{\theta} = \max(X_i)$ has variance of order $\theta^2 / n^2$, much smaller. Let's confirm.
The MLE's variance is $0.000219$, two orders of magnitude below the naive "CRLB" of $0.045$. The actual variance scales as $\theta^2/n^2$, the so-called super-efficient rate. The reason: changing $\theta$ shifts the support, so the score is not defined at the boundary, and the differentiate-then-integrate identity that produces the bound breaks. The Cramér-Rao inequality is silent on this model.
Try it: Consider the shifted exponential $f(x; \theta) = e^{-(x - \theta)}$ for $x \ge \theta$. Is the support free of $\theta$? Sample $n = 100$ at $\theta = 2$ and compute the variance of $\hat{\theta} = \min(X_i)$. Does it scale as $1/n^2$ (super-efficient) or $1/n$ (regular)?
Click to reveal solution
Explanation: The support $[\theta, \infty)$ depends on $\theta$, so the model is non-regular. The variance of $\min(X_i)$ scales as $1/n^2 = 0.0001$, matching our simulation. The CRLB does not apply, and the MLE is super-efficient.
How does the CRLB connect to MLE asymptotic efficiency?
For regular models the maximum likelihood estimator is asymptotically efficient: as $n \to \infty$, $\sqrt{n}(\hat{\theta}_{\text{MLE}} - \theta) \xrightarrow{d} \mathcal{N}(0, I(\theta)^{-1})$. In words, the MLE achieves the Cramér-Rao floor in the limit, even when no finite-sample efficient estimator exists. This is why MLE is the default tool of classical inference: for large $n$ it is essentially optimal, and software packages estimate $I(\theta)$ from the observed Hessian to produce standard errors.
For Poisson, $\bar{X}$ is exactly the MLE and is finite-sample efficient, efficiency is $1$ at every sample size. For models where the MLE is biased in finite samples (Gamma, Beta, log-normal), efficiency starts below $1$ and climbs to $1$ as $n$ grows. That convergence is the asymptotic-efficiency theorem in action.
A practical payoff: every Wald confidence interval you have ever computed comes from inverting the Fisher information matrix at the MLE. For the Normal($\mu, \sigma^2$) example we built earlier, the joint $95\%$ Wald CI is just $\hat{\boldsymbol{\theta}} \pm 1.96 \cdot \sqrt{\operatorname{diag}([n I(\theta)]^{-1})}$.
The CI for $\mu$ covers the true value $2$ comfortably, and the CI for $\sigma^2$ covers $1$. Both intervals were built from the inverse Hessian alone, no closed-form variance formula needed. This pattern generalizes to any well-behaved likelihood: fit by optim() with hessian = TRUE, invert, and you have asymptotic Cramér-Rao-efficient inference for free.
Try it: Build a $95\%$ Wald CI for the Bernoulli parameter $p$ from a sample of $400$ observations at $p = 0.4$, using the observed Fisher information from optim()'s Hessian.
Click to reveal solution
Explanation: Observed Fisher info is the Hessian of the negative log-likelihood at the MLE. Its inverse square root is the asymptotic SE. The CI covers the true $p = 0.4$ as expected.
Practice Exercises
These capstone exercises tie the score, Fisher matrix, and attainment ideas together. Use distinct variable names prefixed with my_ to avoid clobbering tutorial state.
Exercise 1: Build a reusable Fisher information matrix helper
Write fisher_info_matrix(neg_loglik, theta_hat, data) that returns the per-observation Fisher matrix using optim()'s Hessian divided by $n$. Test it on Normal($\mu, \sigma^2$) data with $n = 800$ at $(\mu, \sigma^2) = (1, 4)$ and confirm the diagonal entries match $\operatorname{diag}(1/4,\ 1/32)$.
Click to reveal solution
Explanation: Diagonal entries $0.247$ and $0.0309$ match the closed-form $1/\sigma^2 = 0.25$ and $1/(2\sigma^4) = 0.03125$ to within sampling noise. The off-diagonal is near zero, as expected for the Normal model.
Exercise 2: Asymptotic efficiency of the Gamma MLE
For Gamma(shape = $2$, rate = $1.5$), simulate the MLE of (shape, rate) across $B = 1500$ replications for $n \in \{50, 200, 1000\}$. Use fisher_info_matrix() from Exercise 1 to compute the CRLB at each $n$. Verify that the empirical variance of $\hat{k}$ converges to the (1,1) entry of $[n I(\theta)]^{-1}$.
Click to reveal solution
Explanation: Efficiency climbs from $0.83$ at $n = 50$ to $0.99$ at $n = 1000$, demonstrating asymptotic efficiency of the Gamma MLE. Finite-sample bias drags variance up, but the bias washes out at rate $1/n$.
Exercise 3: Super-efficiency of the bias-corrected Uniform MLE
For Uniform($0, \theta$) with $\theta = 5, n = 100$, the bias-corrected MLE is $\hat{\theta}_c = (n+1)/n \cdot \max(X)$. Simulate $5000$ replicates, compute the variance of $\hat{\theta}_c$, and compute its ratio to the (invalid) "CRLB" $\theta^2/n$. Confirm the ratio is far below $1$.
Click to reveal solution
Explanation: The bias-corrected MLE has variance $0.0026$, ratio $0.01$ relative to the naive "CRLB" of $0.25$. The CRLB is invalid here because the support depends on $\theta$, and the true variance scales as $\theta^2/n^2$ rather than $\theta^2/n$.
Putting It All Together: Complete Inference for the Beta Distribution
Let's run the full Cramér-Rao toolkit on a non-trivial multi-parameter model. We sample from Beta$(\alpha, \beta)$ with $\alpha = 2, \beta = 5$, fit by maximum likelihood, extract the Fisher information matrix from the Hessian, build a covariance estimate, and report Wald confidence intervals for both parameters.
Both $95\%$ Wald confidence intervals cover the true values $\alpha = 2$ and $\beta = 5$. The widths reflect the curvature of the log-likelihood at the MLE, sharper curvature, narrower CI, exactly as the Fisher information formalism predicts. The same optim() + solve(hessian) pattern works for Gamma, Weibull, von Mises, and any other smooth parametric model with regular likelihood.
Summary
- The Cramér-Rao bound is a variance floor: $\operatorname{Var}(\hat{\theta}) \ge 1/[n I(\theta)]$ for scalar $\theta$, generalizing to $[n I(\boldsymbol{\theta})]^{-1}$ in matrix form.
- The bound flows from two facts about the score: it has mean $0$ and variance $I(\theta)$. Cauchy-Schwarz does the rest.
- Attainment is restricted to one-parameter exponential families with a sufficient statistic linear in the score.
- Three regularity conditions are required: support free of $\theta$, twice-differentiable log-likelihood, and the swap of derivative and expectation. Uniform$(0, \theta)$ violates all three.
- The maximum likelihood estimator is asymptotically efficient under regularity: its variance approaches the CRLB as $n \to \infty$.
- In R, the Hessian from
optim(..., hessian = TRUE)is the observed Fisher information; its inverse is the Cramér-Rao-based covariance matrix used for Wald inference.
| Concept | Formula | R idiom |
|---|---|---|
| Score | $U(\theta) = \partial \ell / \partial \theta$ | D(loglik_expr, "theta") |
| Fisher info (scalar) | $I(\theta) = -E[\partial^2 \ell / \partial \theta^2]$ | optim(..., hessian = TRUE)$hessian / n |
| Fisher info matrix | $-E[\nabla^2 \ell]$ | optim(..., method = "BFGS", hessian = TRUE) |
| CRLB (matrix) | $[n I(\boldsymbol{\theta})]^{-1}$ | solve(fit$hessian) |
| Asymptotic SE | $1/\sqrt{n \hat{I}}$ | sqrt(diag(solve(fit$hessian))) |
References
- Lehmann, E. L. & Casella, G., Theory of Point Estimation (2nd ed., Springer 1998), Chapter 2 (Information inequality and Cramér-Rao bound). Link
- Casella, G. & Berger, R. L., Statistical Inference (2nd ed., Duxbury 2002), Section 7.3. Link
- Cox, D. R. & Hinkley, D. V., Theoretical Statistics (Chapman & Hall 1974), Chapter 8.
- Wikipedia, Cramér-Rao bound. Link
- Stanford Stats 200, Lecture 15: Fisher information and the Cramer-Rao bound. Link
- R Core Team,
optim()documentation. Link - Gilbert, P. & Varadhan, R.,
numDerivpackage documentation. Link
Continue Learning
- Sufficient Statistics in R, the Fisher-Neyman factorization theorem and how sufficient statistics relate to attainment of the CRLB.
- UMVUE in R: Rao-Blackwell & Lehmann-Scheffé, turning any unbiased estimator into the minimum-variance unbiased estimator using sufficiency and completeness.
- Exponential Family Distributions in R, the canonical form that guarantees CRLB attainment and connects sufficiency, Fisher information, and conjugate priors.