Chi-Square Goodness-of-Fit Test in R: Does Your Data Fit a Distribution?
The chi-square goodness-of-fit test in R checks whether observed category counts match an expected distribution. You hand chisq.test() your counts and the probabilities you expect, and it returns a single p-value telling you how surprised you should be by the data you saw.
What does the chi-square goodness-of-fit test actually answer?
Suppose you roll a die 60 times. A fair die should land on each face about 10 times. If face 5 came up 14 times and face 2 only 5 times, is the die loaded, or is that just luck? The chi-square goodness-of-fit test answers exactly this question. Pass chisq.test() your counts, and it scores how far the observed counts stray from the uniform distribution you'd expect.
The chi-square statistic is 5.6 across 5 degrees of freedom, and the p-value is 0.35. Because the p-value is well above 0.05, you do not reject the null hypothesis that the die is fair. In plain language: the wobble between counts is the kind you'd see by chance from a fair die roughly one time in three. By default, chisq.test() assumes a uniform distribution, which is why you didn't have to tell it what "fair" means. We'll override that default in the next section.
Try it: You flip a coin 100 times and record 58 heads and 42 tails. Use chisq.test() to test whether the coin is fair. Save the result to ex_coin_test.
Click to reveal solution
Explanation: With only two categories, the default uniform expectation is 50/50. A 58/42 split produces X-squared = 2.56 and p = 0.11, so you can't reject fairness at any standard threshold.
How do you test against custom probabilities?
Real distributions are rarely uniform. Mendel's classic dihybrid pea cross predicts a 9:3:3:1 ratio of yellow-round, yellow-wrinkled, green-round, and green-wrinkled peas. To check whether a sample matches that ratio, you pass the expected probabilities through the p = argument. Mendel's actual 1865 counts for those four categories were 315, 108, 101, and 32. Let's see how close they came.
The p-value of 0.93 is enormous. The observed counts are almost suspiciously close to Mendel's prediction. You can keep the same peas_test object around for the rest of this post, since we'll mine it for residuals and effect size later. Notice that peas_exp had to sum to 1, which is why we divided the 9, 3, 3, 1 weights by 16. R will throw an error if your probabilities don't sum to 1.
chisq.test() requires p to sum to 1. If you have ratios like 9:3:3:1, divide by their total before passing to p, or set rescale.p = TRUE to let R do it for you.Try it: Your marketing team expects website traffic to split 40% organic, 30% paid, 20% social, and 10% email. Last month you saw 412, 285, 220, and 83 visits in those channels. Test whether the actual mix matches the expected mix.
Click to reveal solution
Explanation: A p-value of 0.81 means the observed mix is statistically indistinguishable from the planned mix. The campaign is hitting its targets.
How does R compute the chi-square statistic?
The function reports a number, but where does it come from? Walking through the math by hand makes the output stop feeling like magic. The recipe is simple: for each category, compute how far the observed count strays from the expected count, square it, divide by the expected count, then sum across categories.
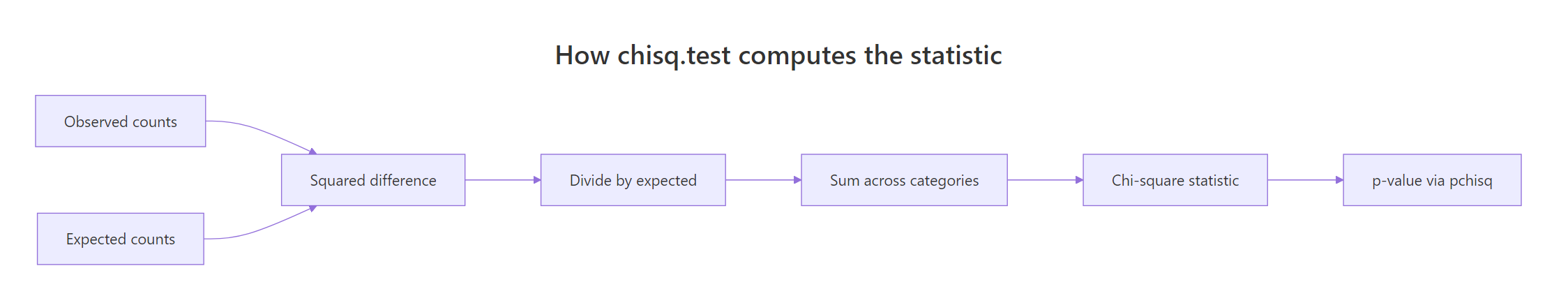
Figure 1: How the chi-square statistic is built from observed and expected counts.
The intuition is that big categories naturally show bigger absolute deviations, so we scale the squared gap by what was expected. That keeps the contribution of each category fair regardless of size. Here is the formula:
$$\chi^2 = \sum_{i=1}^{k} \frac{(O_i - E_i)^2}{E_i}$$
Where:
- $\chi^2$ = the chi-square statistic
- $k$ = the number of categories
- $O_i$ = the observed count in category $i$
- $E_i$ = the expected count in category $i$ ($E_i = N \cdot p_i$, where $N$ is the total sample size)
The degrees of freedom are $k - 1$ for a goodness-of-fit test. The p-value is the area under a chi-square distribution with $k - 1$ degrees of freedom that lies to the right of the observed statistic. Now let's reproduce R's result by hand for the peas data.
The hand-computed values match chisq.test() exactly. The expected counts (312.75, 104.25, 104.25, 34.75) are nearly identical to Mendel's observed counts (315, 108, 101, 32), so each squared deviation is small and the sum stays near zero. The function pchisq() returns the cumulative probability up to a value, so 1 - pchisq() gives the right tail, which is the p-value for goodness-of-fit.
Try it: A factory expects 50% pass, 35% rework, and 15% scrap on its line. In a sample of 200 parts you observe 95 pass, 75 rework, and 30 scrap. Compute the chi-square statistic by hand and compare to chisq.test().
Click to reveal solution
Explanation: Both the manual sum and chisq.test() produce X-squared = 1.07. They have to, because R is doing the same arithmetic.
What assumptions must your data meet?
The chi-square test rests on three assumptions you should verify before trusting the p-value. First, observations must be independent: each die roll, customer, or pea pod is its own data point. Second, categories must be mutually exclusive and exhaustive: every observation falls into exactly one category. Third, expected counts must be at least 5 in every category. The third assumption is the one R will warn you about, because the chi-square distribution approximation breaks down when expected counts get small.
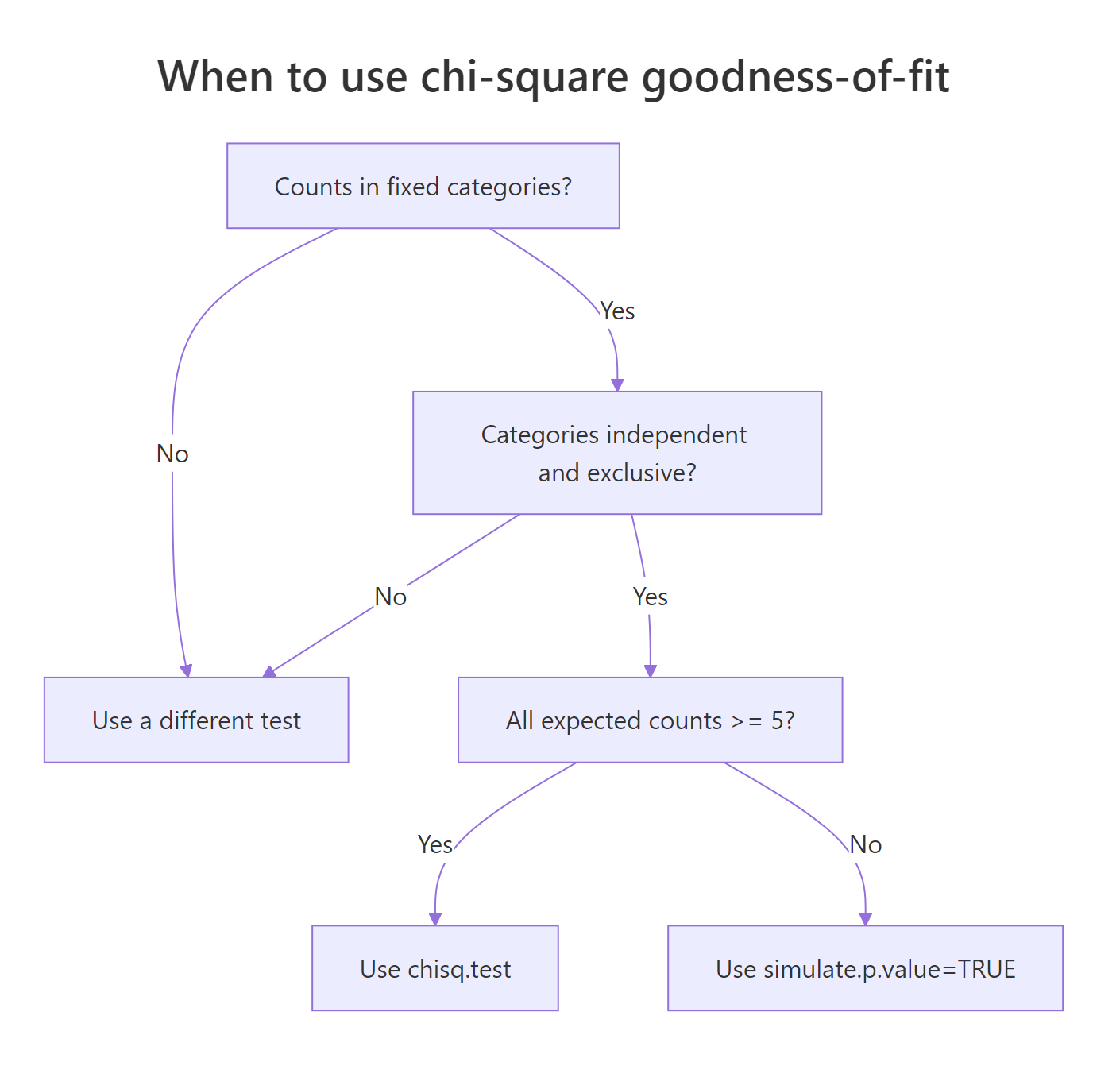
Figure 2: Decision flow: which form of the test to use based on expected counts.
Let's deliberately trigger the small-N warning so you know what it looks like. The trick is to give chisq.test() a sample where one or more expected counts fall below 5.
Every expected count is 3, so all five categories violate the rule. R's warning is real, not cosmetic: with expected counts this small, the reported p-value can be off by a meaningful margin. The fix is to ask R to compute the p-value via Monte Carlo simulation rather than the chi-square approximation. The simulate.p.value = TRUE flag does exactly that, and B controls how many random tables to simulate.
The Monte Carlo p-value is 0.46, which agrees with the asymptotic value here, but you don't get a "may be incorrect" warning. The trade-off is that the test no longer reports degrees of freedom (it shows df = NA) because Monte Carlo doesn't use the chi-square distribution.
simulate.p.value = TRUE to switch to a Monte Carlo p-value.Try it: A pet shop says 1 in 6 of its incoming kittens is black. In a small batch of 12 kittens, 5 are black. Use simulate.p.value = TRUE to test whether the shop's claim is consistent with this batch.
Click to reveal solution
Explanation: Five black kittens in 12 is a lot more than the 2 you'd expect if 1 in 6 were black. The simulated p-value of about 0.013 says this is genuinely unusual, so the shop's claim looks shaky for this batch.
Which categories drive a significant result?
A small p-value tells you that something is off, but not where. To find the culprit categories, look at the standardized residuals. Each category gets a residual that behaves like a z-score: roughly normal with mean 0 and standard deviation 1 under the null. Values larger than about 2 in absolute size flag a category whose count surprised the test most.
Every standardized residual is well below 1 in absolute size, which lines up with the very high p-value. No single category is pulling on the test. For an even quicker read, plot the residuals against horizontal reference lines at +/-2.
All four bars are short and far inside the dashed reference lines. Visually that's the same story the table told: no category broke the test. When you see bars crossing the dashed lines, those categories are the ones to highlight in your write-up.
$residuals returns Pearson residuals: $(O - E) / \sqrt{E}$. $stdres returns standardized residuals, which divide further by an additional variance correction. The standardized version is what you compare to +/-2 because it actually behaves like a z-score under the null.Try it: Customer arrivals across 5 weekdays were 18, 20, 24, 22, 36. Test uniformity, then identify which day has the most surprising count using standardized residuals.
Click to reveal solution
Explanation: Friday's standardized residual is 3.0, well past +/-2, while every other day sits inside the band. The chi-square statistic is large because of Friday alone.
How big is the effect, not just the p-value?
Statistical significance and practical importance are different things. With a huge sample size, a tiny deviation can be highly significant. Effect size measures the size of the deviation in standardized units, independent of $N$. For chi-square, the standard effect size is Cohen's w. It is the square root of the chi-square statistic divided by the total sample size.
$$w = \sqrt{\frac{\chi^2}{N}}$$
Cohen's interpretation thresholds are 0.1 (small), 0.3 (medium), and 0.5 (large). Always report w alongside the p-value so readers can tell whether a "significant" finding is worth acting on.
A w of 0.02 is well below "small" on Cohen's scale, confirming what the p-value already told us: Mendel's data fit the 9:3:3:1 prediction almost perfectly. In a study where you found a "significant" p-value of 0.04 on n = 100,000, a tiny w like 0.02 would let you tell stakeholders "yes it's significant, but the effect is not meaningful."
Try it: A test reports chi-square = 18.4 on a sample of N = 200. Compute Cohen's w and classify it.
Click to reveal solution
Explanation: w = 0.30 sits right at Cohen's "medium" threshold. The deviation from the expected distribution is moderate, neither trivial nor large.
Practice Exercises
These capstone exercises combine multiple concepts from the tutorial. Use distinct variable names so they don't overwrite the tutorial state.
Exercise 1: Customer arrivals over a workweek
A coffee shop expects equal arrivals across Monday through Friday. Last week the counts were 42, 38, 51, 47, 92. Test uniformity, identify the surprising day from standardized residuals, and compute Cohen's w to judge effect size.
Click to reveal solution
Explanation: The p-value is tiny, day 5 (Friday) has a standardized residual of 3.67 (well past +/-2), and Cohen's w of 0.37 is "medium-to-large." Friday is materially busier than the other days, so the manager should staff up.
Exercise 2: Birth months versus month-length probabilities
If birthdays were spread uniformly across the year, every month would get a share equal to its length divided by 365. In a sample of 600 people, the monthly counts (Jan to Dec) were 48, 41, 55, 49, 52, 47, 56, 51, 50, 53, 49, 49. Test the data against month-length probabilities and report the effect size.
Click to reveal solution
Explanation: A p-value of 0.96 and w = 0.085 (below "small") means the observed births are perfectly consistent with month-length proportions. There is no birth-month clustering in this sample.
Putting It All Together
Let's run the full pipeline on a realistic example. A bag of M&M candies is supposed to follow Mars' published color ratio: 24% blue, 20% orange, 16% green, 14% yellow, 13% red, 13% brown. You count one bag and find 32 blue, 22 orange, 20 green, 18 yellow, 14 red, and 14 brown. Does this bag match the company's claim?
Conclusion you can write up: With X-squared = 0.69, df = 5, p = 0.98, w = 0.08, no expected count below 5, and every standardized residual well inside +/-2, the bag's color distribution is statistically indistinguishable from the company's published ratio.
Summary
The chi-square goodness-of-fit test reduces to four moves: get your counts, declare the expected probabilities, run chisq.test(), then probe the result for assumptions, residuals, and effect size.
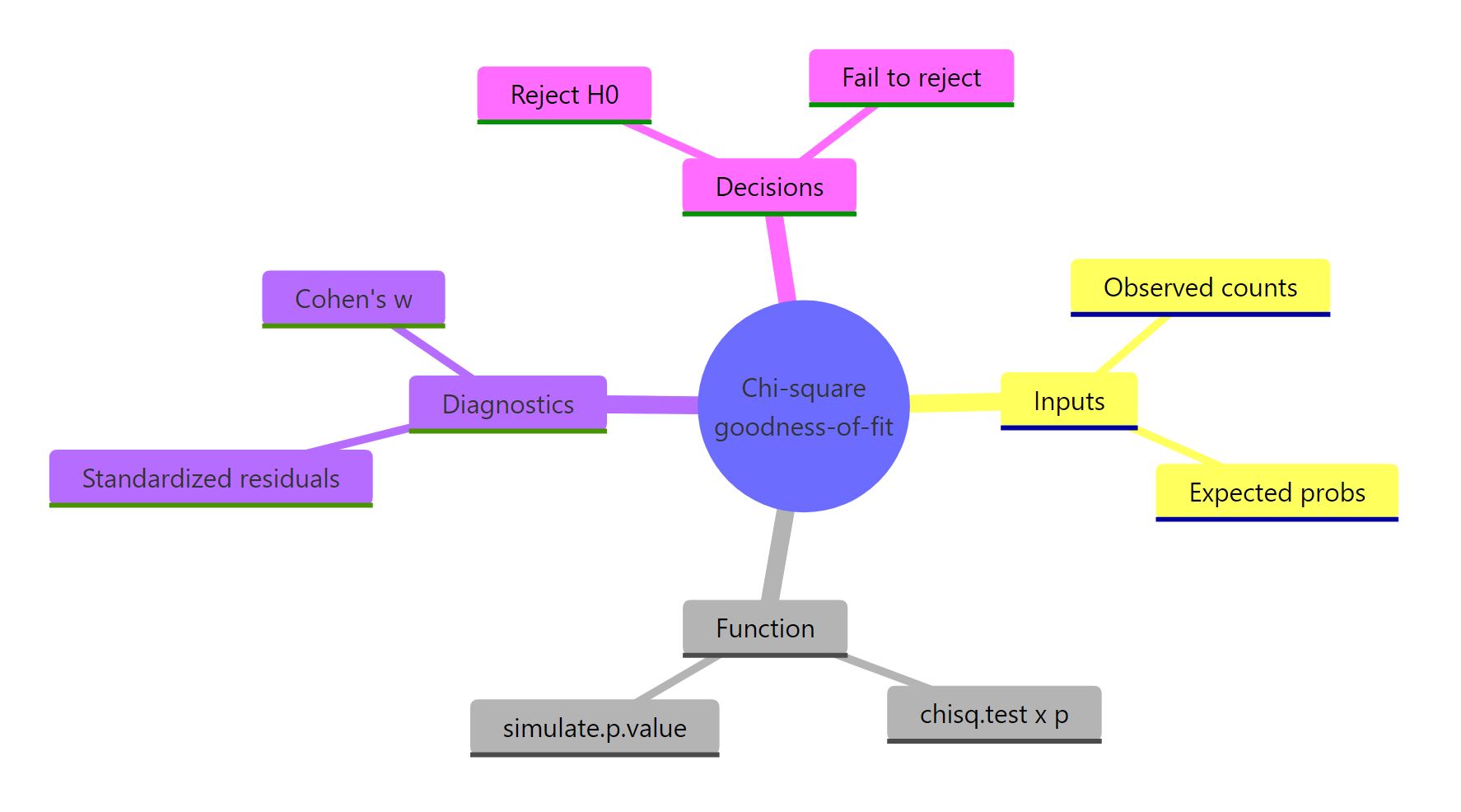
Figure 3: The four moving parts of a goodness-of-fit analysis at a glance.
| Step | Function or attribute | What it returns | When to use |
|---|---|---|---|
| Run the test | chisq.test(x, p) |
Chi-square statistic, df, p-value | Every analysis |
| Check assumptions | result$expected |
Expected counts per category | Before trusting the p-value |
| Handle small N | simulate.p.value = TRUE |
Monte Carlo p-value | When any expected count < 5 |
| Find the culprit | result$stdres |
Z-like residual per category | After a small p-value |
| Quantify effect | sqrt(chi-square / N) |
Cohen's w | Always, alongside the p-value |
| Visualize | ggplot() of residuals |
Bar chart with +/-2 reference | Communicating findings |
References
- R Core Team. chisq.test, base R documentation. Link
- Agresti, A. Categorical Data Analysis, 3rd Edition. Wiley (2013).
- Cohen, J. Statistical Power Analysis for the Behavioral Sciences, 2nd Edition. Routledge (1988). Chapter 7 covers Cohen's w.
- Sharpe, D. "Chi-Square Test is Statistically Significant: Now What?". Practical Assessment, Research & Evaluation, 20(8) (2015). Link
- NIST/SEMATECH. e-Handbook of Statistical Methods, Section 1.3.5.15: Chi-Square Goodness-of-Fit. Link
- Mendel, G. "Versuche über Pflanzenhybriden" (1865), translated as "Experiments in Plant Hybridization."
- Wickham, H. & Grolemund, G. R for Data Science. Chapter on Exploratory Data Analysis. Link
Continue Learning
- Chi-Square Test of Independence in R, the sister test for two-variable association in a contingency table.
- Categorical Data in R, foundations of frequency tables and mosaic plots before you reach for chi-square.
- Chi-Square Tests in R, a tour of the full chi-square family, including independence, homogeneity, and goodness-of-fit.