Exponential Family Distributions in R: Sufficient Statistics & Canonical Form
An exponential family distribution writes its density in a single canonical form: a base measure times an exponential of a natural parameter dotted with a sufficient statistic, minus a log-partition function. Almost every distribution you use in R, Normal, Bernoulli, Poisson, Gamma, belongs to this family, and that shared structure is the engine behind generalized linear models. This tutorial walks the canonical form, sufficient statistics, and the natural-parameter map using base R so everything runs in your browser.
What is an exponential family distribution?
The canonical density of an exponential family distribution has four moving parts: a base measure $h(x)$, a natural parameter $\eta$, a sufficient statistic $T(x)$, and a log-partition function $A(\eta)$. When you plug the right pieces in, a familiar distribution like Normal($\mu$, 1) drops out. Let's verify that by computing the canonical form by hand and comparing it to R's dnorm().
The two columns match to five decimals across all four points, which is the entire claim of the canonical form: every Normal density can be rebuilt from the four pieces above. Notice that $\mu$ enters only through $\eta T(x) - A(\eta)$, never through $h(x)$. That clean separation is what makes the canonical form so useful: parameters live in one chunk and the data lives in another.
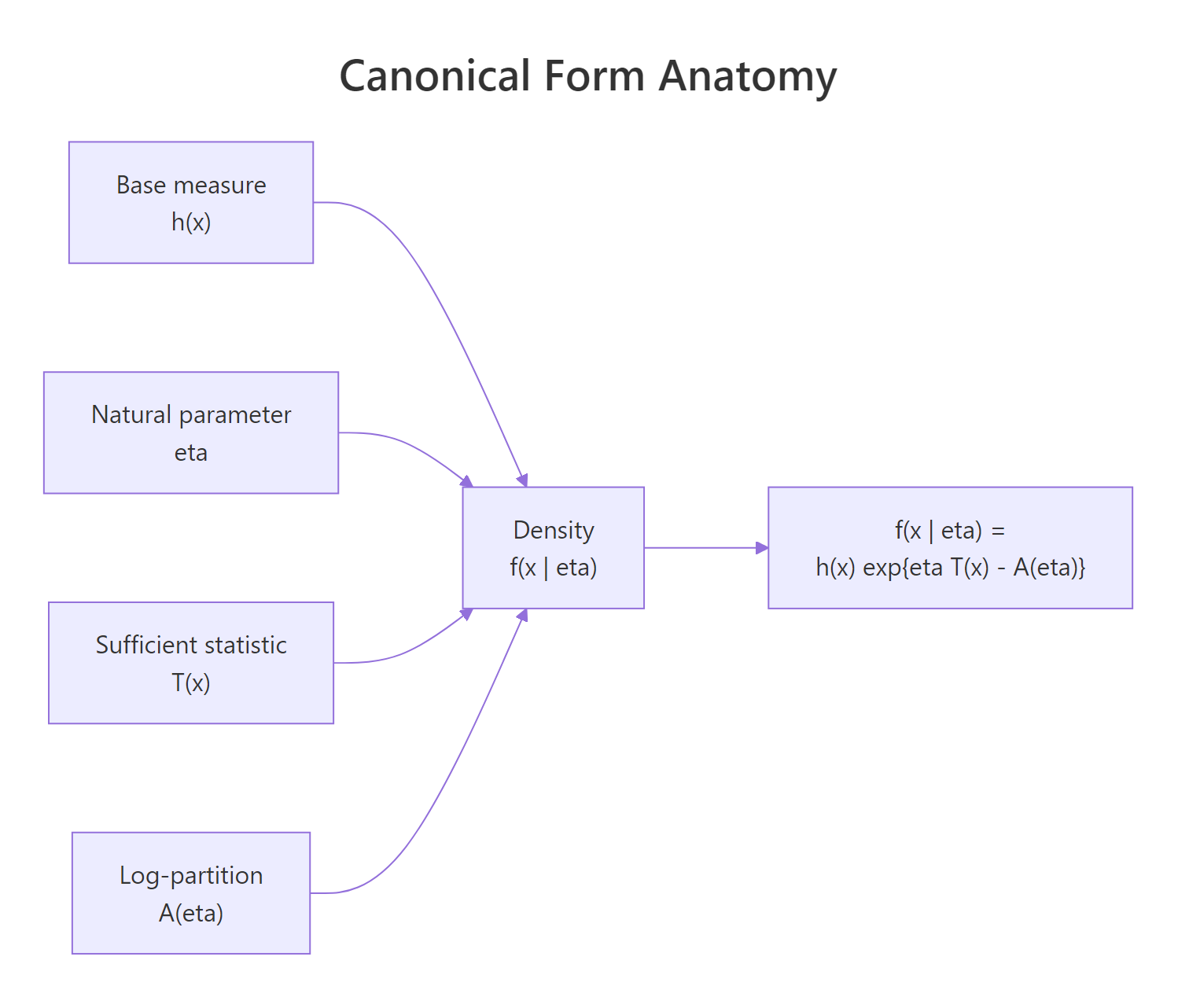
Figure 1: The four building blocks of an exponential family density.
The full density formula is
$$f(x \mid \eta) = h(x) \, \exp\!\left\{ \eta\, T(x) - A(\eta) \right\}$$
Where:
- $h(x)$ is the base measure, the part of the density that does not involve the parameter.
- $\eta$ is the natural parameter (sometimes called the canonical parameter).
- $T(x)$ is the sufficient statistic, a function of the data.
- $A(\eta)$ is the log-partition function (also called the cumulant function), chosen so the density integrates to 1.
If you have ever written $\text{Normal}(\mu, 1)$ as $\frac{1}{\sqrt{2\pi}} \exp\!\left(-\frac{(x - \mu)^2}{2}\right)$ and expanded the square, you have already done the canonical-form rewrite. The trick is recognising that $\mu x$ becomes $\eta T(x)$ and $\mu^2 / 2$ becomes $A(\eta)$.
Try it: Compute the base measure $h(2.5)$ for the standard Normal canonical form by hand and verify it numerically. Use the formula $h(x) = (1 / \sqrt{2\pi}) \exp(-x^2 / 2)$.
Click to reveal solution
Explanation: $h(x)$ is the standard-Normal density's "shape" before any parameter shift. At $x = 2.5$ it is small because 2.5 sits in the right tail.
How do you write Normal, Bernoulli, and Poisson in canonical form?
Three of the most common distributions in R, the Normal, the Bernoulli, and the Poisson, each rewrite into the canonical form with a different choice of $\eta$, $T(x)$, $A(\eta)$, and $h(x)$. Once you see the three side by side, the pattern is hard to unsee. Let's package them as a helper and confirm each canonical density matches its built-in counterpart.
The three families share the same skeleton, only the four pieces change. For Normal, $\eta$ is just $\mu$; for Bernoulli, $\eta$ is the logit of $p$; for Poisson, $\eta$ is the log of $\lambda$. Those transformations look familiar because they are exactly the link functions you meet in glm(). We will return to that connection in the GLM section.
Now let's confirm the canonical form reproduces the textbook density for Bernoulli and Poisson.
Both columns match to four or five decimals, so the canonical form holds. The Bernoulli case is especially clean: the canonical form expresses "probability of success" in log-odds units, which is why logistic regression's natural scale is the logit and not the probability itself. The Poisson case carries the same lesson with log taking the role of logit.
Try it: Show that the Exponential($\lambda$) distribution is in the exponential family by filling in $\eta$, $T(x)$, and $A(\eta)$ for $f(x \mid \lambda) = \lambda \exp(-\lambda x)$, $x > 0$. Compute its canonical density at $x = 1$, $\lambda = 0.5$ and confirm it matches dexp().
Click to reveal solution
Explanation: Pull $\lambda$ out of the density into an exponential: $\lambda e^{-\lambda x} = e^{-\lambda x + \log \lambda}$. The negative sign on $-\lambda x$ becomes $\eta T(x)$ with $\eta = -\lambda$, and $\log \lambda$ becomes $-A(\eta)$ so $A(\eta) = -\log(-\eta)$.
What is a sufficient statistic and why is it useful?
A statistic $T(X_1, \ldots, X_n)$ is sufficient for a parameter $\theta$ if, once you know $T$, the original sample carries no extra information about $\theta$. The technical statement is the Fisher–Neyman factorization theorem: $T$ is sufficient if and only if the joint density factors as $g(T(x); \theta) \cdot h(x)$. For an exponential family, that factorization is built in, the canonical form already separates parameter-dependent and data-dependent pieces, and $T(x)$ is the bridge.
In practice this matters because it means you can throw away the raw data and keep only $T$ without losing any information for inference. For Bernoulli, that "compression" is the count of successes. For Poisson, it is the sum of events. For Normal($\mu$, 1), it is the sample sum.
The maximum-likelihood estimates fall out for free. For a Bernoulli, the MLE of $p$ is the sample mean; for Poisson, the MLE of $\lambda$ is the sample mean; for Normal($\mu$, 1), the MLE of $\mu$ is the sample mean. All three statements say the same thing in canonical-form language: the MLE is the value of $\theta$ that makes the expected sufficient statistic equal the observed one.
Try it: You observe a Bernoulli sample of 50 trials with 32 successes. Without simulating anything, compute the sufficient statistic and the MLE of $p$.
Click to reveal solution
Explanation: The sufficient statistic for Bernoulli is the count of successes, and the MLE of $p$ is that count divided by $n$. You did not need the individual 0/1 outcomes at all.
How does the natural parameter connect to the mean and variance?
The log-partition function $A(\eta)$ is more than a normalising constant. Its derivatives with respect to $\eta$ generate the moments of the sufficient statistic:
$$\frac{dA(\eta)}{d\eta} = \mathbb{E}[T(X)], \qquad \frac{d^2 A(\eta)}{d\eta^2} = \mathrm{Var}(T(X))$$
That is why $A$ is also called the cumulant function: its first derivative is the mean of $T$ and its second derivative is the variance. This identity gives you a free moment computer for any exponential family, no integrals required, and it is what lets glm() do its magic so cheaply.
Let's check the identity for Poisson($\lambda = 2$). We have $A(\eta) = e^\eta$, so $A'(\eta) = e^\eta$ and $A''(\eta) = e^\eta$, which evaluates to 2 at $\eta = \log 2$. That matches the textbook fact that a Poisson's mean equals its variance equals $\lambda$.
Both numerical derivatives recover the value 2 to four decimals, confirming the identity. The same trick works for any member of the family: pick $A(\eta)$, differentiate once to get the mean, differentiate twice to get the variance. For Bernoulli, $A(\eta) = \log(1 + e^\eta)$, so $A'(\eta) = e^\eta / (1 + e^\eta) = p$, the Bernoulli mean.
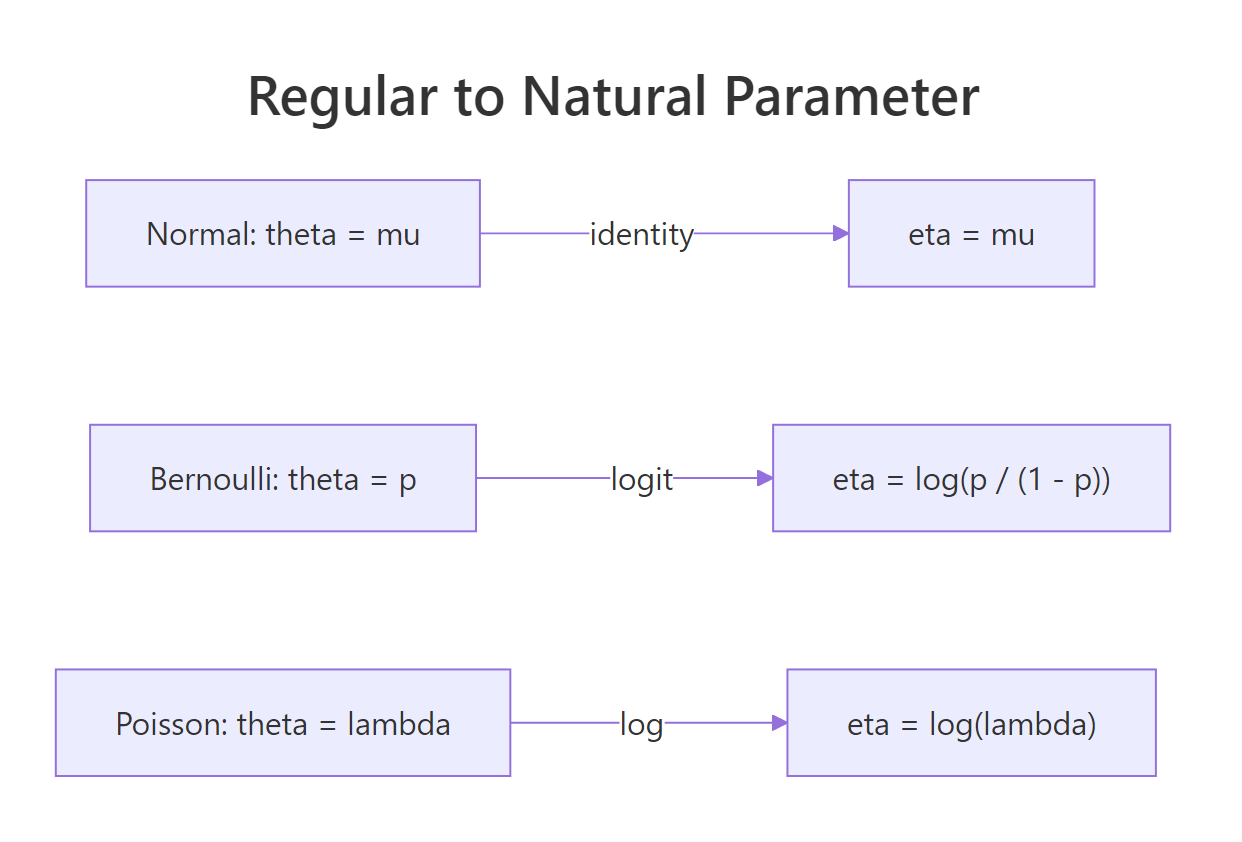
Figure 2: The natural-parameter map for Normal, Bernoulli, and Poisson.
The natural-parameter map itself, $\theta \to \eta(\theta)$, is the canonical link in GLM language: identity for Normal, logit for Bernoulli, log for Poisson. We will use that link directly in the next section.
Try it: Use the cumulant identity to compute the variance of a Bernoulli($p$) random variable from $A(\eta) = \log(1 + e^\eta)$ at $p = 0.3$. You should recover $p(1 - p) = 0.21$.
Click to reveal solution
Explanation: A second numerical derivative of $A(\eta) = \log(1 + e^\eta)$ at $\eta = \mathrm{logit}(0.3)$ recovers $p(1 - p) = 0.21$, the Bernoulli variance.
Why does the exponential family power GLMs?
A generalized linear model writes the conditional distribution of $Y \mid X$ as an exponential family with the natural parameter linked to a linear predictor: $\eta = X\beta$. When the link function is exactly the natural-parameter map (logit for Bernoulli, log for Poisson, identity for Normal), it is called the canonical link. The canonical link is the default in glm() and gives the cleanest score equations, so MLE solves quickly.
Let's fit a Poisson regression and read off the coefficients on the natural-parameter scale.
The fitted equation is $\log \lambda = 0.65 + 0.45 x$, so $\lambda = e^{0.65 + 0.45 x}$, and at $x = 3$ that gives roughly 7.4 events per unit, matching the trend in the small dataset. The crucial thing is that glm() did not "decide" to use a log link, it inherited it from the fact that the Poisson's natural parameter is $\log \lambda$. Same algebra, fancier wrapper.
glm(family = poisson(link = "identity")) for a Poisson with an identity link, but the MLE no longer has a closed-form score equation and convergence often suffers. Stick with canonical links unless a specific scientific reason argues against them.Try it: Without running any code, name the canonical link function for each of these GLM families: Bernoulli, Poisson, Normal($\mu$, 1).
Click to reveal solution
- Bernoulli: logit ($\eta = \log(p / (1 - p))$)
- Poisson: log ($\eta = \log \lambda$)
- Normal: identity ($\eta = \mu$)
Explanation: Each canonical link is exactly the natural-parameter map $\eta(\theta)$ from the canonical form. That is why they are "canonical".
Practice Exercises
Exercise 1: Cast a Geometric distribution as an exponential family
The Geometric distribution counts trials until the first success: $P(X = k) = (1 - p)^{k - 1} p$ for $k = 1, 2, \ldots$. Rewrite this in canonical form by identifying $\eta$, $T(x)$, $A(\eta)$, and $h(x)$, then verify your formula at $p = 0.4$ and $k = 3$ matches dgeom(2, 0.4). Note that R's dgeom() parametrises by the number of failures before success, so subtract 1.
Click to reveal solution
Explanation: The canonical decomposition is not unique. You can absorb scale factors into either $h(x)$ or $A(\eta)$ as long as the product is unchanged. Here we tucked the constant $1 / (1 - p)$ into $h$, giving a clean $A(\eta) = -\log p$.
Exercise 2: Derive the Poisson MLE from the sufficient statistic and verify with glm()
Given a Poisson sample, derive the MLE of $\lambda$ in two ways: (a) from the sufficient statistic by setting the sample mean equal to $\lambda$, and (b) by fitting an intercept-only glm() with the canonical log link and exponentiating the intercept. Confirm the two estimates agree to within numerical tolerance.
Click to reveal solution
Explanation: Both routes agree because the canonical-link GLM solves precisely the score equation "expected sufficient statistic equals observed sufficient statistic", which is the same equation method-of-moments solves. Closed form versus iteratively-reweighted least squares are two roads to the same answer for the canonical case.
Putting It All Together: a Gamma example end-to-end
The Gamma distribution is a two-parameter exponential family with sufficient statistic $T(x) = (\log x, x)$ and natural parameter $\eta = (\alpha - 1, -\beta)$, where $\alpha$ is the shape and $\beta$ is the rate. Let's simulate a Gamma sample, compute the sufficient statistics, and fit the parameters by maximum likelihood using only base R.
The MLE recovers $\hat\alpha \approx 3.05$ and $\hat\beta \approx 0.51$, very close to the true values $\alpha = 3$, $\beta = 0.5$. The sufficient statistics $T_\text{gamma}$ are everything optim() needed to find the MLE: had we given it only the two summary numbers and the sample size, the answer would be the same. This is the "compression for free" promise of the exponential family in action.
Summary
The canonical form gives every member of the exponential family a common structure, and that structure is what powers MLE, GLMs, and most of the closed-form moment formulas you have used.
| Family | $h(x)$ | $\eta$ | $T(x)$ | $A(\eta)$ | Canonical link |
|---|---|---|---|---|---|
| Normal($\mu$, 1) | $\frac{1}{\sqrt{2\pi}} e^{-x^2/2}$ | $\mu$ | $x$ | $\eta^2 / 2$ | identity |
| Bernoulli($p$) | $1$ | $\log\!\frac{p}{1-p}$ | $x$ | $\log(1 + e^\eta)$ | logit |
| Poisson($\lambda$) | $1 / x!$ | $\log \lambda$ | $x$ | $e^\eta$ | log |
| Exponential($\lambda$) | $1$ | $-\lambda$ | $x$ | $-\log(-\eta)$ | reciprocal |
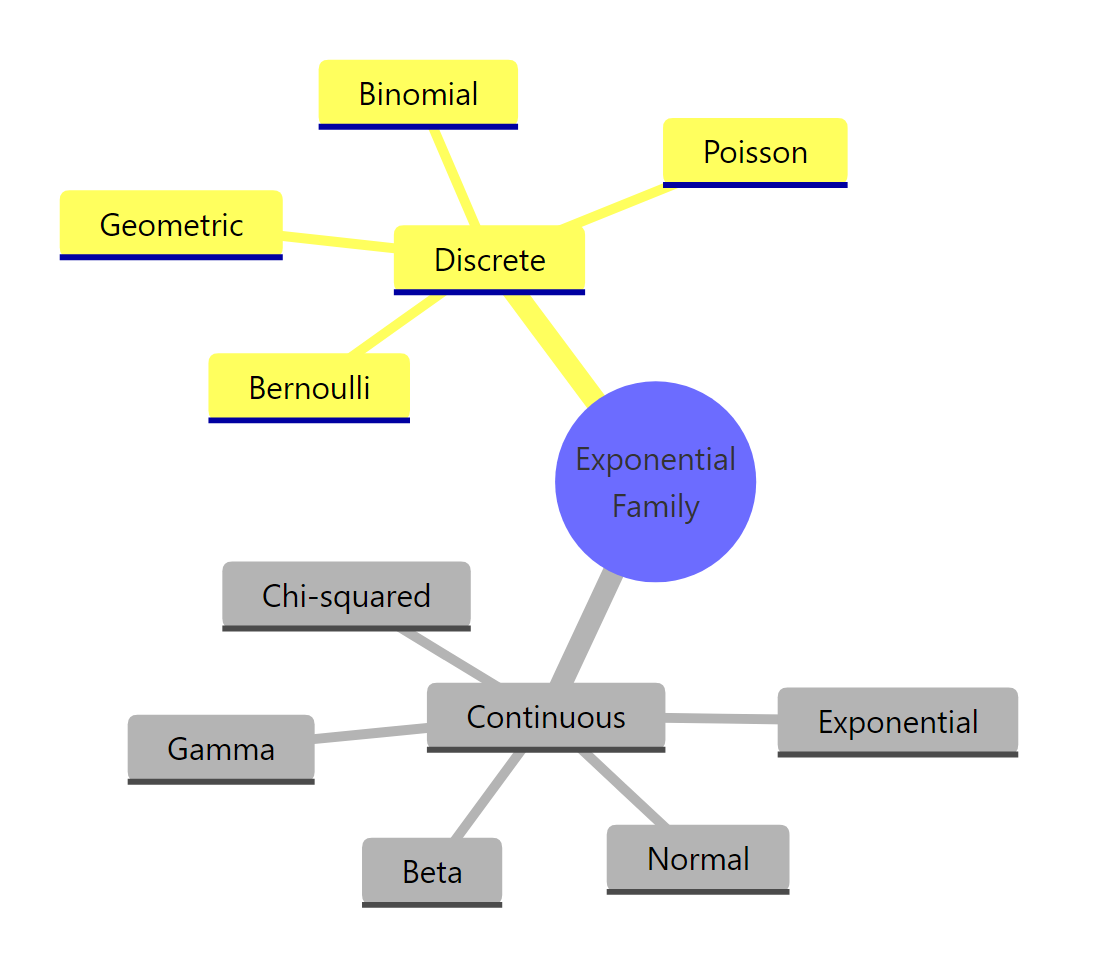
Figure 3: Common distributions that belong to the exponential family.
Key takeaways:
- The canonical density factors data and parameter cleanly into $h(x) \exp\{\eta T(x) - A(\eta)\}$.
- The sufficient statistic $T(x)$ is everything the data tell you about $\theta$, so MLE depends only on $T$ and $n$.
- The natural-parameter map $\theta \to \eta$ is the GLM canonical link, identity for Normal, logit for Bernoulli, log for Poisson.
- Differentiating the log-partition function once gives the mean of $T$, twice gives the variance.
References
- Casella, G., & Berger, R. L. Statistical Inference, 2nd Edition. Duxbury (2002). Chapter 3.4: The exponential family. Link
- Wikipedia. Exponential family. Link
- DasGupta, A. The Exponential Family and Statistical Applications, Purdue University lecture notes. Link
- Geyer, C. J. Stat 5421 Lecture Notes: Exponential Families, University of Minnesota. Link
- Jordan, M. I. An Introduction to Probabilistic Graphical Models, Chapter 8: The exponential family. UC Berkeley. Link
- R Core Team. The R Stats Package: glm(). Link
- McCullagh, P., & Nelder, J. A. Generalized Linear Models, 2nd Edition. Chapman & Hall (1989). The original GLM textbook.
Continue Learning
- Maximum Likelihood Estimation in R, how the score equation behind the canonical-link GLM works in detail.
- Logistic Regression with R, the Bernoulli exponential family at work, with the logit link in front.
- Poisson Regression in R, count-data modelling using the Poisson canonical form.