Mann-Whitney U Test in R: Two Independent Groups, Distribution-Free
The Mann-Whitney U test compares two independent groups when the data are not normal or are ordinal. In R, wilcox.test(y ~ group) returns the U statistic and a p-value without assuming any distribution.
How do you run a Mann-Whitney U test in R?
Suppose you want to know whether automatic and manual cars get different miles per gallon, but a few outliers are pulling the means around. The Mann-Whitney U test ranks every observation across both groups and compares the rank totals, so a single odd value cannot dominate the result. One call to wilcox.test() does the whole thing.
The U statistic (R prints it as W) is 42, and the p-value is 0.001871. With p well below 0.05, you reject the null that automatic and manual cars come from the same distribution: manual cars get systematically more miles per gallon. That is your answer in three lines of code, no normality assumed.
Try it: Use wilcox.test() to compare Sepal.Length between the setosa and versicolor species in the iris dataset. Save the result to ex_iris_mw and print it.
Click to reveal solution
Explanation: Setosa flowers have far shorter sepals than versicolor, so the rank sums diverge and the p-value is essentially zero.
When should you use Mann-Whitney over a t-test?
The two-sample t-test asks whether two group means differ, and it trusts the Central Limit Theorem to keep the math honest. That trust falls apart with skewed data, ordinal scales, or small samples with outliers. The Mann-Whitney U test side-steps the problem by working on ranks, which compress big values and pull in the tails.
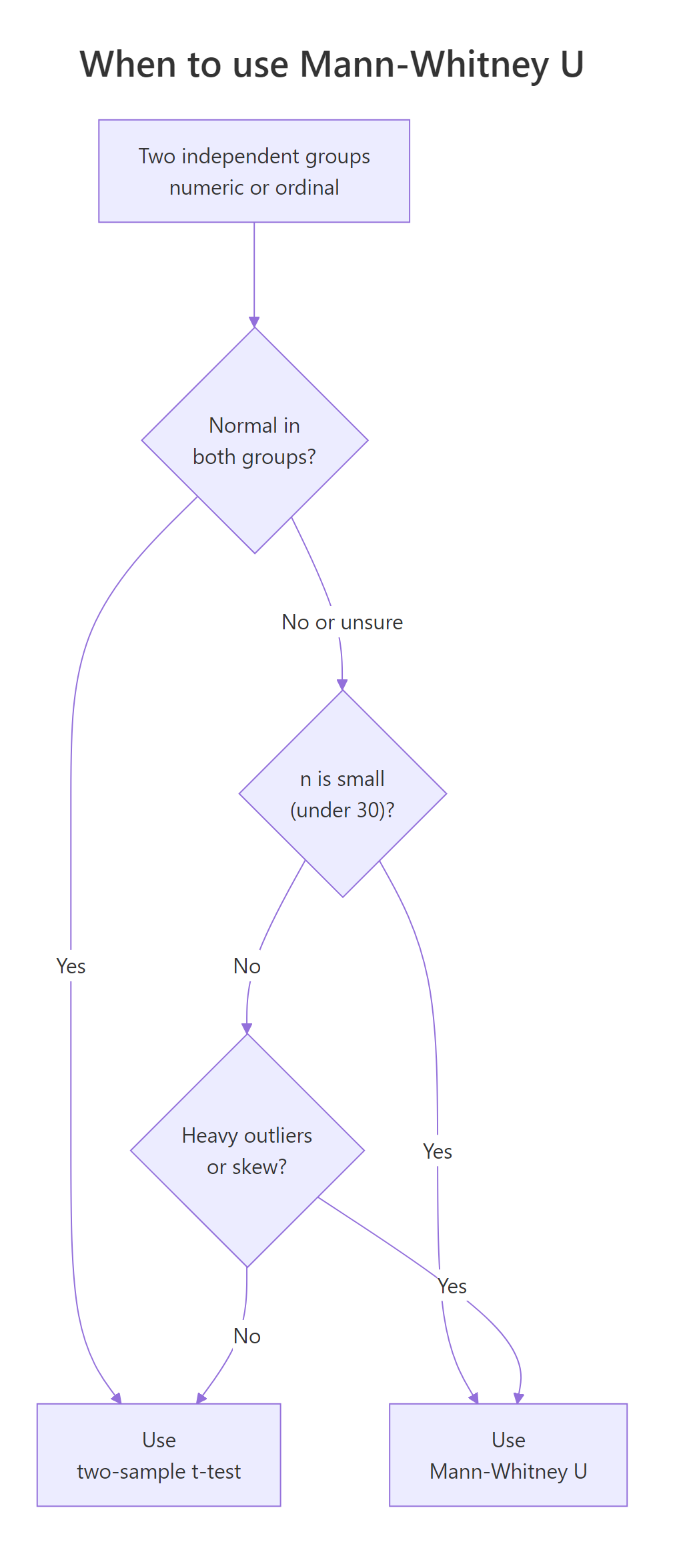
Figure 1: When to choose Mann-Whitney U over a two-sample t-test.
Let's see what happens when the t-test's normality assumption is wrong. We will simulate two heavily right-skewed samples and run both tests on the same data.
Same data, two different verdicts. The t-test gives p = 0.1010 (not significant), while the Mann-Whitney returns p = 0.0479 (significant at 5%). The skew inflated the t-test's variance estimate and washed out the signal; ranks did not care.
Try it: You collect a 5-point Likert satisfaction score (1 = bad, 5 = great) from two product variants, n = 12 each. Which test fits the data, and why? Set your answer as a string in ex_choice.
Click to reveal solution
Explanation: Likert ratings have ranked spacing but unequal psychological distances between points, so a t-test on the raw integers misrepresents the data.
How do you run the test from a data frame using formulas?
Splitting a column into two vectors before testing is fine for two groups, but it gets tedious fast. R's formula syntax keeps the data frame intact and reads almost like a sentence: outcome ~ group. The result is identical to the vector form, but the code is cleaner and ports straight to other modelling functions.
The formula mpg ~ am says "model mpg by am." Inside wilcox.test(), that becomes "compare mpg between the two levels of am." The data = mtcars argument tells R where to find both columns.
Same W = 42, same p = 0.001871. The formula form costs one line and gives you free integration with subset = filters and dplyr pipelines. Use it whenever your data already lives in a tidy data frame.
wilcox.test(mpg ~ am, data = mtcars, subset = cyl != 8) runs the test on 4- and 6-cylinder cars only, no preprocessing needed. The vector form would force a separate filter step.Try it: Use the chickwts dataset (chicken weights by feed type). Run a Mann-Whitney U test comparing casein and horsebean feeds using formula syntax and store it in ex_chick_mw.
Click to reveal solution
Explanation: subset = feed %in% c(...) keeps only the two levels you want. Casein-fed chicks weigh far more, so the p-value is tiny.
How do you visualize the two groups before testing?
A test gives you a number. A picture tells you what that number means. Before you trust any p-value, plot the two groups so you can see the shape, the spread, and any outliers. If the two distributions look wildly different in shape, the test still runs, but your interpretation must change.
We will use ggplot2 to draw a boxplot of mpg by am, with am cast as a factor so it reads as a category instead of a number.
The manual box sits clearly above the automatic box, and the spreads are roughly comparable. That visual is what makes the p = 0.001871 from earlier feel right rather than mysterious.
A density plot adds detail the boxplot hides, namely the shape of each group and any hint of bimodality.
Both curves are roughly symmetric, the manual curve is shifted right, and the shapes are similar. With matching shapes you can interpret the Mann-Whitney result as "manual cars have a higher median mpg." If the shapes had differed, you would phrase it more cautiously as "manual cars tend to have higher mpg."
Try it: Add a violin plot layer on top of the boxplot for mpg by am. Save the plot to ex_violin.
Click to reveal solution
Explanation: geom_violin() shows the full density, and a narrow geom_boxplot() overlay keeps the median and quartiles visible.
How do you measure effect size for Mann-Whitney?
A p-value tells you whether a difference is real. An effect size tells you whether it is worth caring about. With large samples, even tiny differences become statistically significant; without an effect size, you cannot tell a meaningful gap from a trivial one.
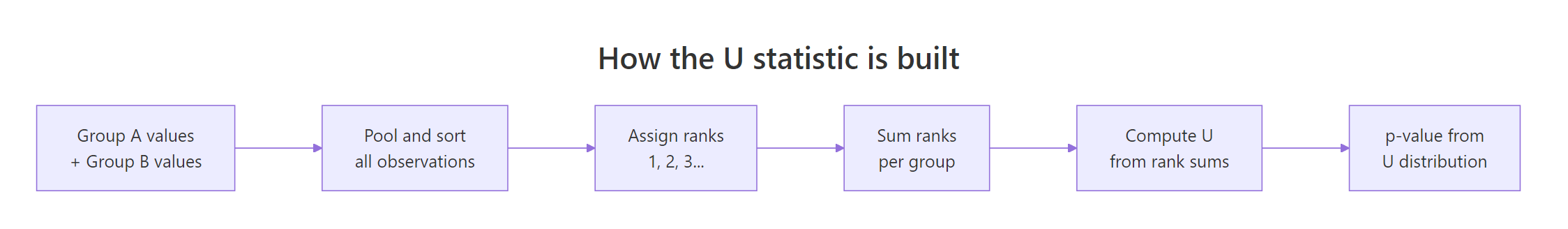

Figure 2: How the U statistic is built from ranks across both groups.
The natural effect size for the Mann-Whitney test is the rank-biserial correlation, which Kerby (2014) showed equals the simple difference between the proportion of pair comparisons where group A wins and the proportion where group B wins. The Wendt formula computes it directly from the U statistic and the two group sizes.
$$r_{rb} = 1 - \frac{2U}{n_1 \cdot n_2}$$
Where:
- $U$ = the W statistic R prints from
wilcox.test() - $n_1$ = size of the first group passed to the test
- $n_2$ = size of the second group
The rank-biserial correlation is 0.66. The sign tells you the direction: a positive value means group 2 (manual) tends to have larger values, since R orders levels alphabetically (or by factor levels) when it computes U. The magnitude tells you the strength.
Cohen's thresholds for r rate 0.66 as a large effect, which lines up with the visual gap in the boxplot. Always report the effect size with three decimals plus a verbal label so the reader does not need to remember the cutoffs.
Try it: Compute the rank-biserial correlation for the setosa vs versicolor Sepal.Length test from the first exercise. Use ex_iris_mw$statistic. Save it to ex_iris_r.
Click to reveal solution
Explanation: The negative sign means setosa values rank below versicolor values. The magnitude 0.866 is a very large effect.
How do you handle ties, small samples, and one-sided tests?
Three real-world wrinkles trip up beginners: tied values, small samples, and one-sided hypotheses. Each has a one-line fix.
When two or more observations share the same value, R cannot compute the exact p-value and prints a warning. The fix is to switch to the asymptotic approximation with exact = FALSE. The continuity correction stays on by default and adjusts for the discrete-to-continuous jump.
No warning, and the p-value (0.6991) is reliable for any sample where ties are common. The half-rank in W = 15.5 shows that R averaged the ranks for the tied 4s and 7s.
exact = TRUE only when n is small AND there are no ties. Default exact = NULL lets R decide based on sample size and ties. Forcing exact = TRUE with ties produces the warning and falls back to the same approximation anyway.For a directional question, like "is automatic mpg lower than manual mpg?", swap the two-sided default for a one-sided test. The argument names are intuitive once you read them as "the first group's location is less/greater than the second's."
The one-sided p-value (0.0009) is exactly half the two-sided value when the test agrees with the direction. The 95% CI for the location shift, $(-\infty, -3.5)$, says manual cars get at least 3.5 mpg more than automatics with 95% confidence.
Try it: Run a one-sided test on mtcars checking whether mpg_auto is less than mpg_manual, but this time on the data frame using formula syntax. Save it to ex_one_sided.
Click to reveal solution
Explanation: With formula notation, alternative = "less" means level 0 (automatic) has a lower location than level 1 (manual). The p-value matches the vector form to the last digit.
How do you report Mann-Whitney U test results?
A solid report sentence packs the test name, group sizes, statistic, p-value, effect size, and effect direction into one line. Reviewers and readers can verify everything they need without rerunning your code.
The template below uses paste0() so it works in vanilla base R. Plug in your numbers and you have a publication-ready sentence.
That is one sentence, six numbers, zero ambiguity. Drop it straight into a results paragraph.
Try it: Build a similar one-line report for the iris setosa-vs-versicolor result. Save the string to ex_iris_report.
Click to reveal solution
Explanation: Pull each number directly from the test object so you cannot copy-paste-corrupt them.
Practice Exercises
Exercise 1: Compare ozone levels between two months
Using the airquality dataset, test whether Ozone differs between Month == 5 (May) and Month == 8 (August). Handle the missing values, run the test, compute the rank-biserial r, and print a one-line report.
Click to reveal solution
Explanation: August ozone is materially higher than May, with a medium-large negative rank-biserial correlation.
Exercise 2: Chick weight at day 21 by diet
Using ChickWeight, compare weight at Time == 21 between Diet == 1 and Diet == 4. Make a boxplot, run the test with exact = FALSE, compute the effect size, and print a one-line report. Save the test object as my_chick.
Click to reveal solution
Explanation: Diet 4 produces clearly higher day-21 weights, with a large negative rank-biserial correlation.
Exercise 3: Show t-test fragility on skewed data
Simulate two right-skewed samples (n = 25 each) using rexp() with rates 1.0 and 0.6. Run both t.test() and wilcox.test() 500 times with different seeds, count how often each rejects at alpha = 0.05, and compare. Save the two counts as my_t_rej and my_mw_rej.
Click to reveal solution
Explanation: The Mann-Whitney rejects more often than the t-test when the data are skewed, because ranks are robust to long right tails. Your exact counts will vary by a few units.
Complete Example
Let's walk through the full pipeline on the built-in sleep dataset, which records extra hours of sleep for 10 patients on each of two soporific drugs.
The p-value sits just above 0.05, but the rank-biserial r of -0.49 is a medium-large effect. With only 10 per group the test lacks power to crown the difference significant. The honest report is: "we observed a medium effect favouring Drug 2, but n is too small to confirm it." That is exactly the kind of nuance an effect-size-aware reader catches and a p-value-only reader misses.
Summary
| Argument / output | What it does |
|---|---|
wilcox.test(x, y) |
Two-sample test from two vectors |
wilcox.test(y ~ g, data = df) |
Two-sample test from a data frame |
paired = FALSE (default) |
Mann-Whitney U (independent groups) |
alternative = "two.sided" / "less" / "greater" |
Two-sided or directional |
exact = NULL / TRUE / FALSE |
Exact vs asymptotic p-value (must be FALSE with ties) |
conf.int = TRUE |
Adds a CI for the location shift |
correct = TRUE (default) |
Continuity correction for the asymptotic version |
W in output |
Mann-Whitney U statistic |
| Rank-biserial r | Effect size: $1 - 2U / (n_1 n_2)$ |
References
- Mann, H. B., & Whitney, D. R. (1947). On a Test of Whether one of Two Random Variables is Stochastically Larger than the Other. Annals of Mathematical Statistics, 18(1), 50-60. Link
- R Core Team.
wilcox.test()reference, stats package. Link - Conover, W. J. (1999). Practical Nonparametric Statistics, 3rd ed. Wiley.
- Hollander, M., Wolfe, D. A., & Chicken, E. (2014). Nonparametric Statistical Methods, 3rd ed. Wiley.
- Kerby, D. S. (2014). The simple difference formula: An approach to teaching nonparametric correlation. Comprehensive Psychology, 3, 11-IT. Link
- Divine, G. W., Norton, H. J., Barón, A. E., & Juarez-Colunga, E. (2018). The Wilcoxon-Mann-Whitney procedure fails as a test of medians. American Statistician, 72(3), 278-286. Link
- Wickham, H., & Grolemund, G. R for Data Science. Link
Continue Learning
- Wilcoxon Signed-Rank Test in R: the paired counterpart to Mann-Whitney, for matched samples or repeated measures.
- When to Use Nonparametric Tests in R: the decision flowchart that picks Mann-Whitney, Wilcoxon, Kruskal-Wallis, or a parametric alternative for any setup.
- Kruskal-Wallis Test in R: the nonparametric extension to three or more independent groups, plus post-hoc Dunn tests.