Bayes' Theorem in R: Why Medical Tests Mislead You, A Simulation That Shows Why
Bayes' theorem updates the probability of an event given new evidence by combining a prior belief, the likelihood of the evidence under each hypothesis, and the overall probability of the evidence, letting you compute, for example, the chance you actually have a disease given a positive test.
What does Bayes' theorem actually compute?
A 99% accurate test for a disease that strikes 1 in 1,000 people sounds reliable. Yet if you test positive, the chance you actually have the disease is closer to 9% than 99%. This counter-intuitive result is exactly what Bayes' theorem computes. Let's see it in R.
The classical statement of Bayes' theorem is short. Given two events $A$ and $B$:
$$P(A \mid B) = \frac{P(B \mid A) \cdot P(A)}{P(B)}$$
Where:
- $P(A \mid B)$ is the posterior, the probability of $A$ after seeing $B$.
- $P(B \mid A)$ is the likelihood, the probability of seeing $B$ if $A$ is true.
- $P(A)$ is the prior, your belief about $A$ before seeing $B$.
- $P(B)$ is the evidence, the overall probability of $B$ under all hypotheses.
For medical testing, $A$ is "patient has the disease" and $B$ is "patient tested positive." The most useful quantity is $P(\text{disease} \mid \text{positive})$, called the positive predictive value or PPV. Let's wrap the math in a function and call it once.
The output reads "0.0902," meaning that on a positive test for this disease, there is only a 9% chance the patient actually has it. The other 91% are false alarms. The function maps three inputs to a posterior probability, applying Bayes once for each call. Notice how (1 - spec) carries the false positive rate from healthy people, and how it gets weighted by (1 - prevalence), the much larger group.
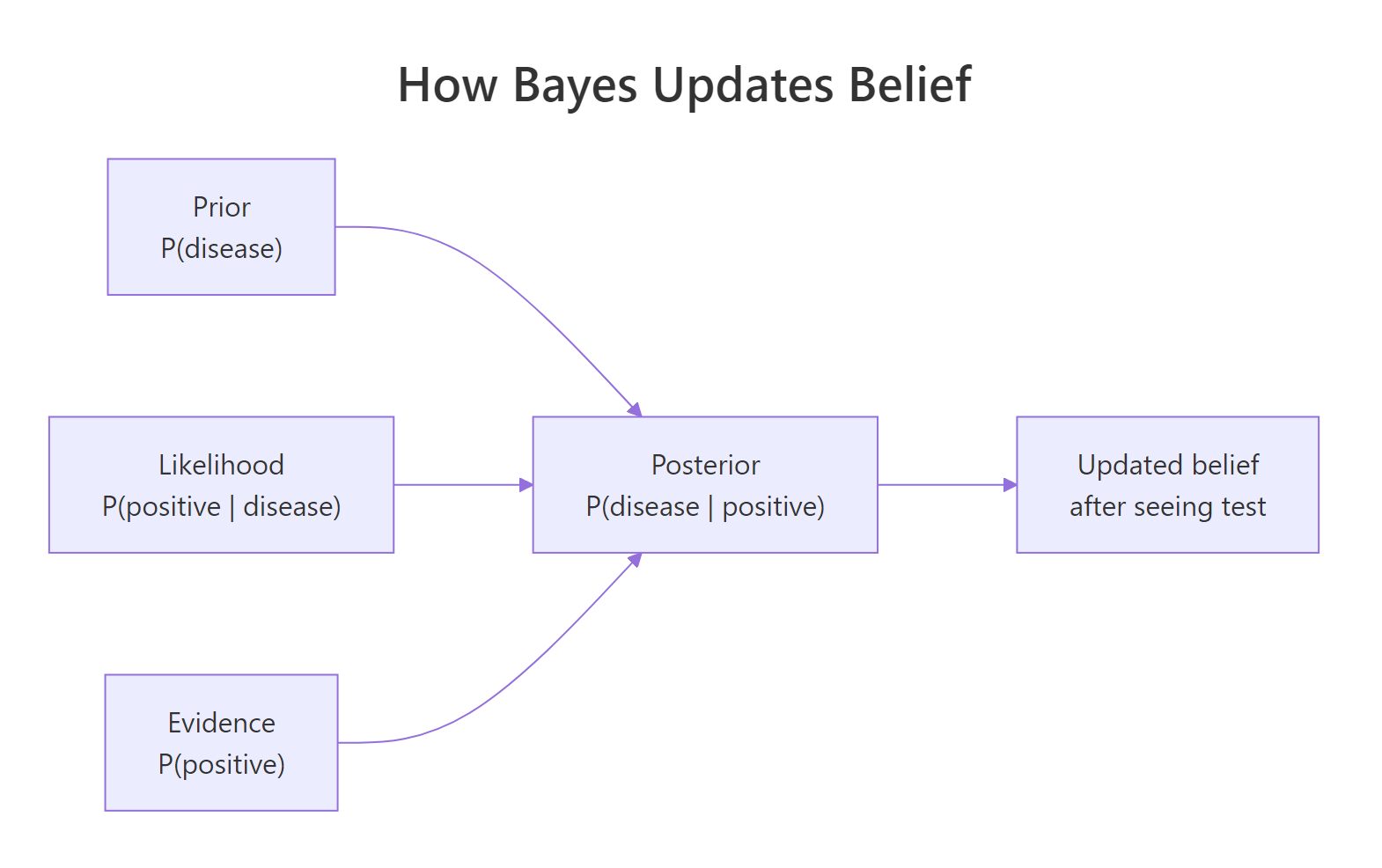
Figure 1: How Bayes combines prior, likelihood, and evidence into a posterior.
To see how PPV moves with disease prevalence, we hold sensitivity and specificity fixed at 99% and sweep the prior across a wide range.
The curve rises steeply from near zero at very low prevalence and approaches 100% only when prevalence is itself high. The same test is "99% accurate" in every scenario, yet its trustworthiness for any single positive result depends entirely on the prior. That single fact is the engine behind every misunderstanding about medical screening.
Try it: Use bayes_pp() to compute the PPV when prevalence is 5%, sensitivity is 95%, and specificity is 90%. Round to two decimal places.
Click to reveal solution
Explanation: At 5% prevalence with 90% specificity, false positives still outnumber true positives nearly 2 to 1, so PPV lands at 33%, not 95%.
Why does a 99% accurate test return mostly false positives?
The answer is arithmetic, not statistics. When the disease is rare, the population of healthy people is enormous, and even a small false positive rate inside that huge group produces more false positives than the entire group of true positives combined.
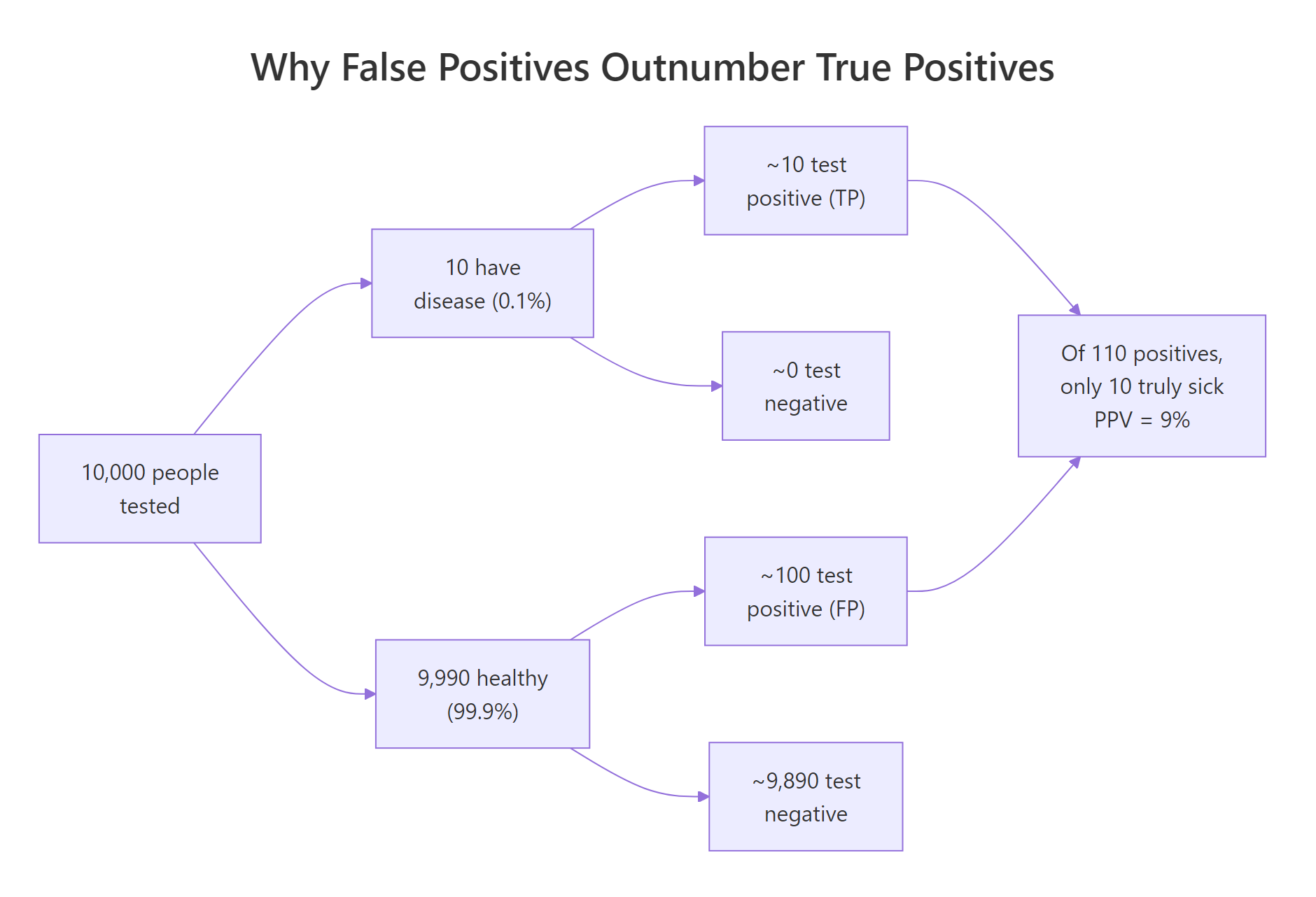
Figure 2: Why false positives can outnumber true positives when a disease is rare.
To make this concrete, let's simulate 10,000 people. We'll set prevalence to 0.1%, draw who has the disease, then assign each person a test result based on sensitivity (for the sick) and specificity (for the healthy). Counting outcomes turns the abstract probability into something you can see.
The table tells the story directly. Among 9,990 healthy people, 100 still tested positive (false positives), while only 10 of the 10 sick people tested positive (true positives). Of the 110 positive results in total, just 10 are real, which gives an empirical PPV of 10 / 110, or about 9%.
The two numbers agree to within sampling noise. The simulation is not telling us anything new beyond what the formula said; it is showing us the same result in counts of people, which our intuition handles better than ratios. The arithmetic of false positives is exactly Bayes' theorem in disguise.
Try it: Re-run the simulation with prevalence <- 0.10 (a 10% community prevalence rather than 0.1%) and observe how the empirical PPV changes.
Click to reveal solution
Explanation: At 10% prevalence the sick group is 100 times larger than at 0.1%, so true positives now outnumber false positives by roughly 11 to 1. PPV jumps from 9% to over 90%, even though the test itself is unchanged.
How do sensitivity, specificity, and prevalence each shift PPV?
Three numbers drive PPV, and they pull in different directions. Sensitivity is $P(\text{positive} \mid \text{disease})$: how often the test catches a real case. Specificity is $P(\text{negative} \mid \text{no disease})$: how often the test correctly clears a healthy person. Prevalence is just $P(\text{disease})$, the prior. To see which lever matters most when the disease is rare, we hold prevalence at 0.1% and sweep specificity from 80% to 99.9%.
The curve is almost flat below 95% specificity and explodes upward above 99.5%. Going from 95% to 99.9% specificity is the difference between a 2% PPV and a 50% PPV. Sensitivity helps you find disease, but specificity is what lets you trust a positive result when the disease is rare.
Try it: Wrap the formula in a tiny helper called ex_ppv() that takes prevalence, sensitivity, and specificity and returns PPV. Call it for prevalence 0.02, sens 0.98, spec 0.95.
Click to reveal solution
Explanation: At 2% prevalence with 95% specificity, only 29% of positives are real. The function reuses the same Bayes formula as bayes_pp() with renamed arguments.
What changes when you test twice?
A second positive test pushes the posterior much higher than a single test, but the math is not "99% accurate twice equals 99.99% accurate." Each test updates the prior with new evidence. Yesterday's posterior becomes today's prior. As long as the two tests are independent, you can apply Bayes twice in a row.
After one positive, PPV is 9%. After a second positive, PPV jumps to 91%. The same test, applied twice, took the result from "almost certainly a false alarm" to "very likely real." This is exactly how confirmatory testing works in clinics: the first test acts as a screen, and a second independent test is required before any diagnosis.
Try it: What is the posterior probability of disease after three consecutive positive tests, all with sensitivity 99% and specificity 99%, starting from a prior of 0.001?
Click to reveal solution
Explanation: Each application multiplies the odds in favour of the disease by the likelihood ratio. After three independent positives at 99% sensitivity and specificity, the posterior approaches certainty.
Where else does Bayes' theorem apply outside medicine?
The medical example is just one application of a general rule. Spam filters use Bayes to compute $P(\text{spam} \mid \text{words})$. Search engines rank pages by $P(\text{relevant} \mid \text{query})$. Machine learning classifiers, naive Bayes most directly, use the same arithmetic on text and image features. The events change, the formula does not.
Given the prior that 40% of emails are spam, seeing the word "lottery" pushes the posterior to 95% spam. The same arithmetic that gave us PPV in medicine gives us the spam probability here. Whether the events are diseases, emails, or stock movements, Bayes is the bookkeeping rule for combining evidence with prior belief.
Try it: Compute the posterior probability of spam given the word "meeting," using the same priors and likelihoods.
Click to reveal solution
Explanation: "Meeting" is much more common in ham than spam, so seeing it lowers the posterior to about 14%, well below the 40% prior. The same formula gives different answers because the likelihoods flip.
Practice Exercises
These three exercises combine the ideas from earlier sections. Use distinct variable names like my_data and my_result so you don't overwrite the tutorial's variables in the same notebook session.
Exercise 1: COVID-style screening test
A community has 2% prevalence of an infection. The available test is 95% sensitive and 98% specific. Compute the PPV of a single positive test. Then find the lowest specificity that pushes PPV above 90% at the same prevalence and sensitivity.
Click to reveal solution
Explanation: At 98% specificity the PPV is only 49%. Pushing specificity to 99.54% raises PPV above 90%. Specificity is the dominant lever when prevalence is low.
Exercise 2: Two independent tests on a population
Simulate 50,000 people with 1% disease prevalence. Each person takes two independent tests, both with sensitivity 95% and specificity 97%. For each scenario below, compute the empirical PPV from the simulation and compare to Bayes:
(a) one positive test, (b) two consecutive positive tests, (c) one positive followed by one negative.
Click to reveal solution
Explanation: A single positive at 1% prevalence gives 25% PPV. Two consecutive positives push it to 91%. A positive followed by a negative collapses it to roughly 1%, near the prior. Each test moves the belief in proportion to its likelihood ratio.
Exercise 3: Naive Bayes email classifier
You have a vocabulary of five words: bonus, meeting, urgent, report, prize. Prior P(spam) = 0.4. Per-class word probabilities are below. Classify a 3-word email containing bonus, urgent, prize by computing the posterior probability of spam, treating the words as conditionally independent.
Click to reveal solution
Explanation: All three words are far more likely under spam than ham, so their likelihood ratios multiply, driving the posterior to over 99%. Naive Bayes scales this exact computation to thousands of words.
Putting It All Together
A clinic deploys two tests for a disease with 1% prevalence. Test A is 99% sensitive and 95% specific (a fast screen). Test B is 90% sensitive and 99.5% specific (a slow confirmatory test). We compute PPV after each test alone and after both tests positive.
The summary makes the policy recommendation obvious. Test A by itself flags far too many false positives to act on (17% PPV), and Test B alone is decent (65%). But Test A followed by Test B both positive lifts the posterior above 97%. This is the rationale for two-stage screening: a sensitive screen catches everyone, then a specific confirmer rules out the false alarms.
Summary
| Takeaway | What it means in code |
|---|---|
| Bayes = prior × likelihood / evidence | bayes_pp(prev, sens, spec) returns the posterior. |
| Low prevalence amplifies false positives | At 0.1% prevalence and 99% specificity, PPV is 9%, not 99%. |
| Specificity dominates PPV when disease is rare | Sweeping specificity reshapes PPV far more than sweeping sensitivity. |
| Sequential tests update the prior | Yesterday's posterior is today's prior; multiply likelihood ratios. |
| Same math powers spam, search, and ML | Naive Bayes is just Bayes' theorem on word features. |
References
- Gigerenzer, G. Reckoning with Risk: Learning to Live with Uncertainty. Penguin, 2002. The original framing of base rate fallacy in medical testing. Link
- McElreath, R. Statistical Rethinking, 2nd Edition. CRC Press, 2020. Chapter 1: The Golem of Prague. Link
- Wickham, H. and Grolemund, G. R for Data Science, 2nd Edition. O'Reilly, 2023. Chapters on simulation and dplyr summaries. Link
- R Core Team. An Introduction to R. CRAN, 2024. Link
- Diagnostic Testing and Bayes' Theorem. Medical Maths blog, 2024. R simulation walkthrough. Link
Continue Learning
- Conjugate Priors in R explains the family of priors that make posteriors exact, no MCMC required, building directly on the belief-update intuition above.
- Grid Approximation in R shows how to compute exact posteriors for any small model by evaluating Bayes on a grid of parameter values.
- MCMC in R introduces Metropolis-Hastings, the algorithm that scales Bayes to high-dimensional models when the integrals stop being tractable.