Regression Tables in R: modelsummary vs stargazer vs gtsummary, Which to Use
A regression table turns a fitted model into a formatted grid of coefficients, standard errors, significance stars, and goodness-of-fit statistics that readers can scan at a glance. In R, three packages dominate this job: modelsummary, stargazer, and gtsummary. This tutorial shows you which one to pick and how to use each.
What makes a regression table publication-ready?
Anyone who has copy-pasted summary(lm(...)) into a Word document knows the feeling. The output is fine for the person who fit the model. It is unreadable for anyone else. A real regression table compresses the same fit into a compact grid where rows are coefficients and columns are either models or estimates. Here is the payoff. One line of R turns a bare lm() into a clean table your co-author will not complain about.
Notice what you got without any formatting work. Coefficients with standard errors stacked below, fit statistics at the bottom, clean column header. That is the baseline that every package in this tutorial aims for, with different defaults and different ways to customise it.
Try it: Fit a one-predictor model on mtcars (ex_fit1 <- lm(mpg ~ wt, data = mtcars)) and print it with modelsummary().
Click to reveal solution
Explanation: modelsummary() accepts any fit object broom::tidy() understands. Single model, single column, same one-line call.
summary() in three ways. They show models side by side, let you pick which statistics appear, and export to the format you actually publish in, HTML, Word, LaTeX, or PDF.How do modelsummary, stargazer, and gtsummary differ at a glance?
The three packages look similar on the surface, but they come from different traditions and optimise for different end products. modelsummary is a generalist built on top of the broom ecosystem. It supports more than a hundred model types out of the box and treats side-by-side comparison as the default. stargazer is the classic LaTeX-first package. It shaped what a "regression table in R" looks like for a decade and is still the default in many economics workflows. gtsummary comes from medical research. Its defaults assume categorical covariates, odds ratios, and clinical reporting conventions.
Here is a side-by-side feature matrix to anchor the rest of the tutorial.
| Feature | modelsummary | stargazer | gtsummary |
|---|---|---|---|
| Default output | tinytable (HTML, LaTeX, Word) | Text, LaTeX, HTML | gt (HTML), flextable (Word) |
| Side-by-side models | Built-in | Built-in | Via tbl_merge() |
| Robust SEs | vcov = "robust" |
Manual with sandwich |
add_vcov() |
| Model types | 100+ via broom/parameters | 40+ hard-coded | Most broom-tidy models |
| Default style | Three-star significance | Journal-style dashes | Clinical, OR/HR columns |
| Active development | Yes (2026) | Maintenance mode | Yes (2026) |
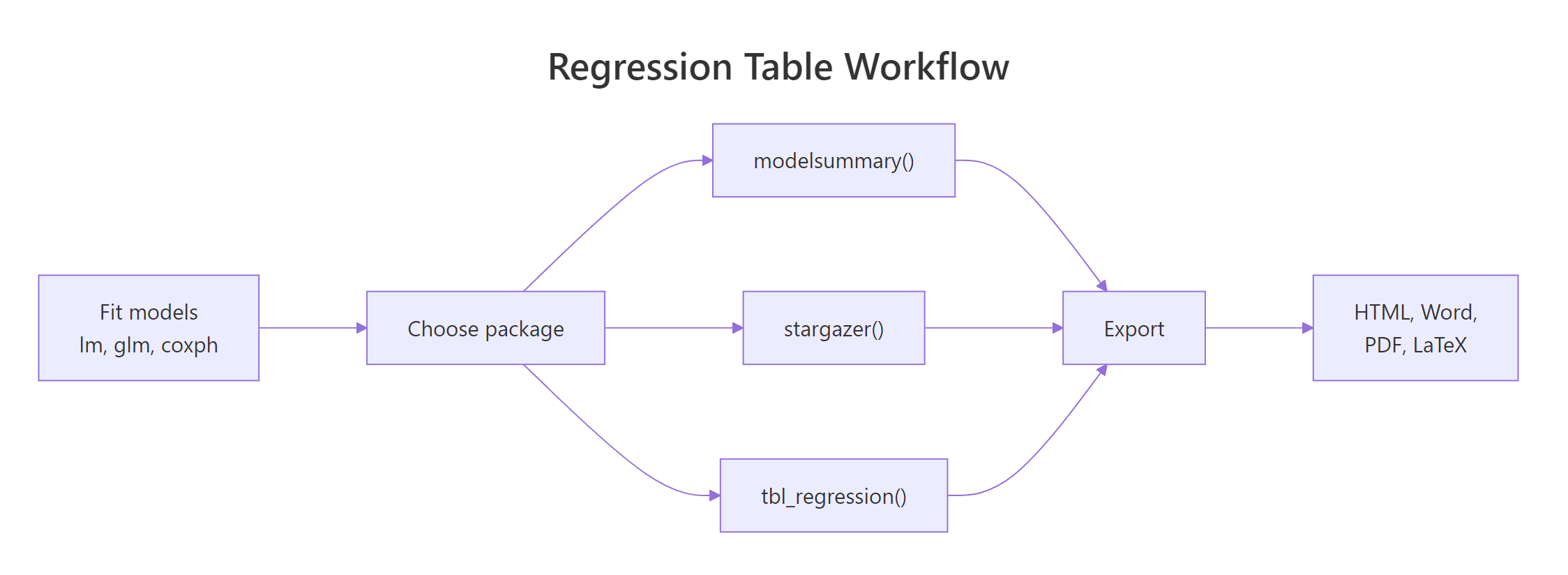
Figure 1: Every package follows the same three-step workflow: fit, table, export.
Try it: Given the feature matrix above, name one scenario where each package is the best choice.
Click to reveal solution
Explanation: gtsummary's built-in defaults for odds ratios, hazard ratios, and categorical reference rows save hours in clinical writing. stargazer and modelsummary both produce journal-ready LaTeX. For quick, multi-model comparisons with minimal code, modelsummary wins.
How do you build the same table with each package?
The fastest way to internalise the differences is to build the same table three ways. You fit the models once, then pass the same list to each package. Each call takes one line.
The named list is the key. Every package in this tutorial reads the names and uses them as column headers, so you get "Base", "+HP", "+Cyl" as column labels instead of "Model 1", "Model 2", "Model 3".
Three models appear as three columns. The stars = TRUE argument adds significance stars with the standard thresholds ( = 0.001, = 0.01, = 0.05). Each coefficient sits above its standard error in parentheses.
Notice the differences. stargazer uses horizontal rules, puts the intercept as "Constant" at the bottom, and adds a "Dependent variable" banner across the top. This is the look of most economics and political science journals.
gtsummary takes a different route. It builds one table per model, then merges them with tbl_merge(). The default shows a 95% confidence interval instead of a standard error, and a p-value column, both clinical conventions.
modelsummary and gtsummary both read list names. list("Base" = fit1, "+HP" = fit2) produces headers "Base" and "+HP"; an unnamed list gives you "Model 1", "Model 2", etc.Try it: Fit ex_fit_iris <- lm(Sepal.Length ~ Petal.Length + Petal.Width, data = iris) and build a stargazer text table for it.
Click to reveal solution
Explanation: type = "text" prints ASCII. Change to "latex" for a journal manuscript or "html" for a webpage.
How do you customize coefficients, standard errors, and fit stats?
Defaults get you a working table. Customisation gets you the table you actually want to ship. All three packages let you rename coefficients, reorder rows, select which fit statistics to show, and swap in robust standard errors. The syntax differs, but the concepts line up one to one.
Four arguments did the heavy lifting. coef_map renames and reorders rows. gof_map picks which fit statistics to show. stars redefines the thresholds. vcov = "robust" uses HC3 heteroskedasticity-robust standard errors, so the SEs in the output differ from the defaults above.
stargazer uses covariate.labels for renaming (in model order, not by name), omit.stat to drop fit statistics, and star.cutoffs to set thresholds. For robust SEs you compute the variance-covariance matrix yourself and pass a list of standard error vectors.
gtsummary uses formula-based labels, a separate estimate_fun argument for numeric formatting, and add_significance_stars() to append stars to the estimate. It leans heavily on the pipe (|>) to chain these steps.
vcov = "robust", "HC1", "HC3", or a function. In gtsummary, pipe through add_vcov(). In stargazer, compute the matrix with sandwich::vcovHC() and pass a list of SE vectors to the se argument, which takes more setup.Try it: Rebuild the modelsummary(models) table with coef_map renaming wt to "Weight" and hp to "HP" only, leaving other rows untouched.
Click to reveal solution
Explanation: When coef_map is partial, only the mapped rows are renamed and reordered. Unmapped rows are dropped unless you include them in the mapping.
How do you export tables for PDF, Word, and HTML?
The piece that separates "fits in my notebook" from "lands in the published paper" is export. Each package is strongest in its home format. modelsummary is the most portable, gtsummary is the cleanest for Word, and stargazer is the fastest for LaTeX.
| Target format | modelsummary | stargazer | gtsummary | |
|---|---|---|---|---|
| HTML | output = "file.html" |
type = "html" |
`as_gt() \ | > gt::gtsave()` |
| LaTeX | output = "file.tex" |
type = "latex" |
as_kable_extra() |
|
| Word (.docx) | output = "file.docx" |
Via HTML + copy-paste | as_flex_table() |
|
| Markdown | output = "markdown" |
type = "text" |
as_kable() |
|
| Quarto / Rmd | Auto-detects | Needs results = "asis" |
Auto-detects |
modelsummary picks the format from the file extension. .docx triggers the flextable backend, .tex triggers LaTeX, .html triggers tinytable's HTML, and .md emits knitr's markdown kable.
gtsummary separates table content from the rendering backend. You build the table once, then convert to gt for HTML (which supports CSS styling) or flextable for Word (which preserves tables cleanly in .docx).
output argument. modelsummary() inspects the rendering target (HTML, PDF, or Word) and picks a sensible default. Hard-coding output = "html" breaks the LaTeX build.Try it: Export the models list to a markdown string and store it in ex_md.
Click to reveal solution
Explanation: output = "markdown" returns the table as a character vector you can paste into a document or log to a file.
Which package should you choose?
Here is the decision framework. Start from your output format and the conventions of your field, not from which package you saw first.
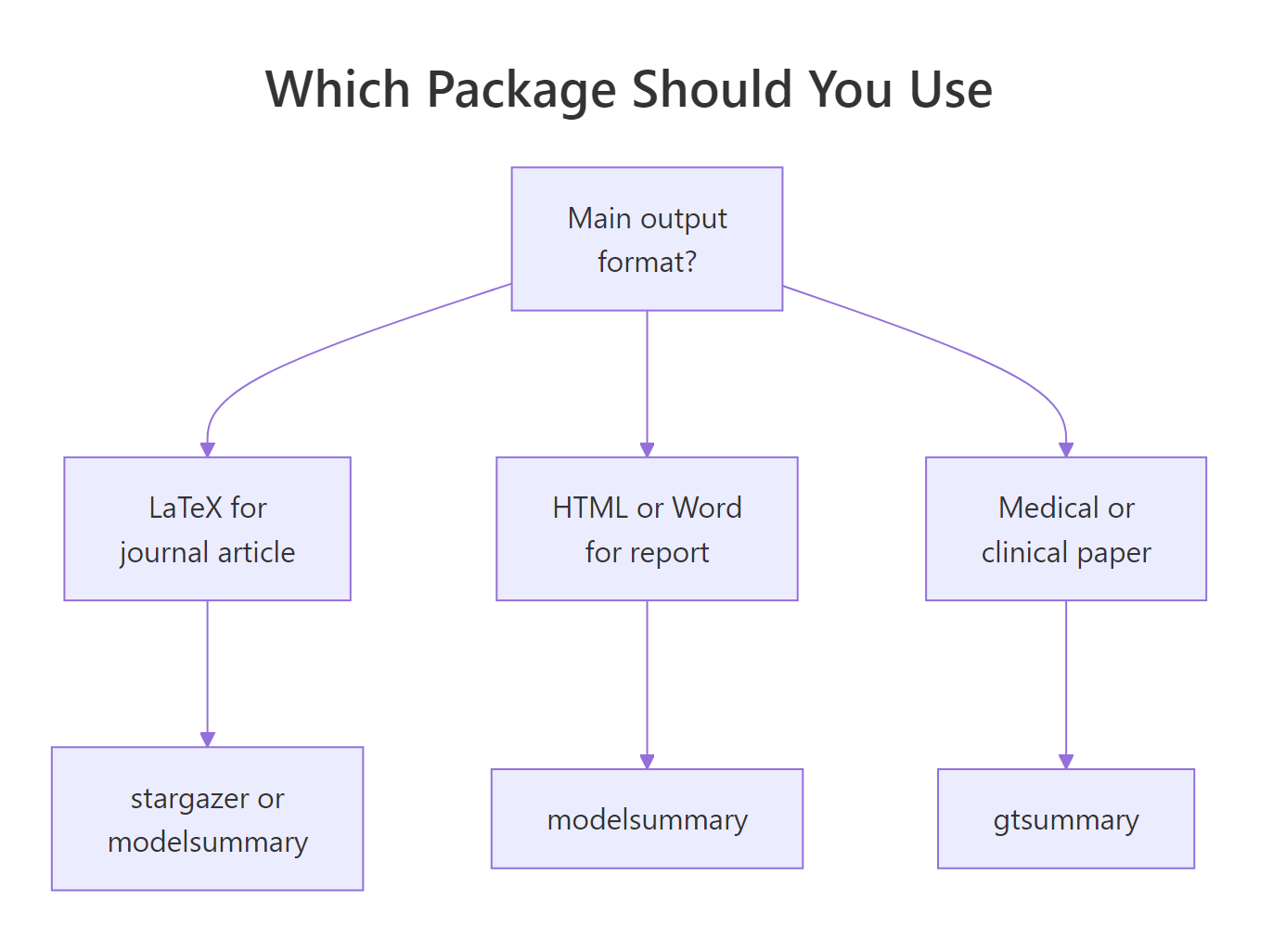

Figure 2: A quick way to pick based on your output format.
The shortlist by use case:
- LaTeX for an economics or finance journal: use
modelsummaryorstargazer. Both produce journal-compatible LaTeX.stargazerwins on default style,modelsummarywins on active development and model coverage. - Word or HTML for a client report: use
modelsummarywithoutput = "file.docx". Single call, portable, round-trips in Microsoft Word without manual fixes. - Clinical or medical paper: use
gtsummary. The defaults assume odds ratios, categorical reference rows, and p-values, which match clinical reporting conventions (CONSORT, STROBE). - Comparing many models side by side: use
modelsummary.gof_map,coef_map, and the named list interface scale cleanly to six or eight columns. - You need a model class stargazer does not support: use
modelsummary. Any object with abroom::tidy()method works, includingfixest,lme4,brms,survival, and many more.
modelsummary. It is the most generalist and future-proof of the three. Reach for gtsummary when you publish medical research. Keep using stargazer only if you already have a working pipeline; it is in maintenance mode and new R model classes may not render cleanly.Try it: For a fit from coxph(Surv(time, status) ~ age + sex), which package gives publication-ready output with the least code?
Click to reveal solution
Explanation: gtsummary recognises coxph objects automatically and exponentiates coefficients into hazard ratios by default. The table header says "HR" and reference rows appear for categorical covariates without extra configuration.
Practice Exercises
Exercise 1: Three-model comparison with custom labels
Using mtcars, fit three nested models predicting mpg, store them in a named list called my_models, and build a modelsummary table with coefficient labels renamed, three significance stars, and nobs plus aic as the only goodness-of-fit rows.
Click to reveal solution
Explanation: The named list gives you named columns. coef_map provides rename plus reorder; gof_map selects which fit rows appear.
Exercise 2: Replicate a stargazer table with modelsummary
Given this stargazer call, reproduce the same table with modelsummary.
Click to reveal solution
Explanation: covariate.labels becomes coef_map. dep.var.labels becomes the list name. omit.stat becomes gof_map with only the stats you want to keep.
Exercise 3: Export a gtsummary logistic regression to Word and markdown
Fit a logistic regression predicting am (auto vs. manual transmission) from mpg and hp using glm(). Build a gtsummary table and convert it once to flextable (for Word) and once to as_kable() (for markdown).
Click to reveal solution
Explanation: exponentiate = TRUE turns log odds into odds ratios. The header switches from "Beta" to "OR" automatically. The same table object converts to flextable for Word or kable for markdown without refitting.
Complete Example
Here is the end-to-end workflow most analysts run once per paper: fit the models, customise, add robust SEs, and export. The output is a self-contained HTML file you can email or attach to a pull request.
Change output = "markdown" to "table.html", "table.docx", or "table.tex" and you have the same table in your publication format of choice. No reshaping, no copy-paste into Excel.
Summary
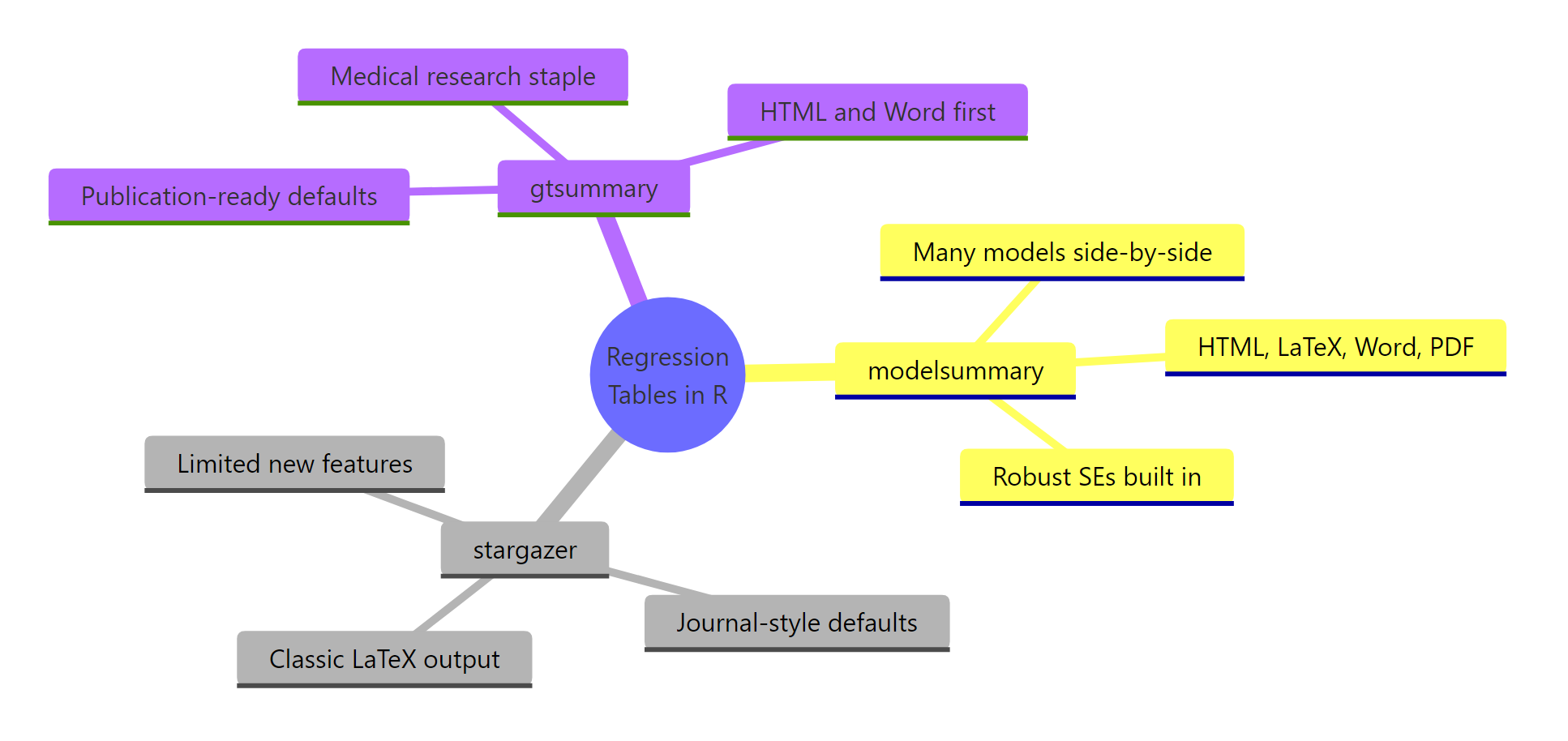
Figure 3: The three packages and their strongest use cases.
| Package | Pick when... | Skip when... |
|---|---|---|
| modelsummary | You want one tool for every workflow, many model types, active development | Your field mandates gtsummary defaults |
| stargazer | Your existing LaTeX pipeline uses it and works | You need a model type added after 2020 |
| gtsummary | You write clinical or medical papers, need OR/HR defaults | You need LaTeX-first journal output |
Key takeaways:
- All three accept the same fit objects, so switching packages never means refitting models.
- Named lists give you clean column headers in modelsummary and gtsummary.
- Robust SEs are one argument in modelsummary (
vcov = "robust"), one pipe in gtsummary (add_vcov()), and manual work in stargazer. modelsummaryis the generalist choice in 2026;gtsummaryis best for medical publishing;stargazerremains usable but is in maintenance mode.
References
- Arel-Bundock, V. (2022). modelsummary: Data and Model Summaries in R. Journal of Statistical Software, 103(1). Link
- modelsummary documentation. Link
- Hlavac, M. (2022). stargazer: Well-Formatted Regression and Summary Statistics Tables. CRAN package version 5.2.3. Link
- Sjoberg, D., Whiting, K., Curry, M., Lavery, J., Larmarange, J. (2021). Reproducible Summary Tables with the gtsummary Package. The R Journal, 13(1). Link
- gtsummary tbl_regression vignette. Link
- Princeton Library, A Hands-on R Tutorial Using Stargazer. Link
- Tilburg Science Hub, Generate Regression Tables in R with the modelsummary Package. Link
- tinytable documentation (modelsummary's default backend since v2.0). Link
Continue Learning
- Linear Regression: fit the lm() models this tutorial formats.
- Logistic Regression with R: the glm() binary-outcome models that gtsummary exponentiates into odds ratios.
- Linear Regression Assumptions in R: verify your fits before you report them in a table.