Chi-Square Test of Independence in R: Assumptions, Effect Size & Power
The chi-square test of independence checks whether two categorical variables are related in a population by comparing the counts you observe in a contingency table to the counts you'd expect if the variables were independent. Most tutorials stop at the p-value; this one walks the full pipeline, assumption checks, effect size with Cramer's V, and a power analysis you can run on your own data.
What does the chi-square test of independence actually answer?
Suppose you have survey data with two categorical columns, say smoking status and exercise level, and you want to know whether they move together or are unrelated. The chi-square test of independence answers exactly this. It compares the counts you actually observed against the counts you would expect if the two variables had nothing to do with each other, then turns the gap into a single p-value. Let's run one now.
You ran the test in two lines. The chi-square statistic is 5.49 with 6 degrees of freedom, and the p-value is 0.48. Because the p-value is well above any standard threshold (0.05, 0.01), you do not reject the null hypothesis of independence. In plain language: smoking status and exercise level look unrelated in this sample.
Try it: Run the same test on survey$Smoke against survey$Sex to check whether smoking varies by sex in this dataset. Save the fitted test to ex_xt.
Click to reveal solution
Explanation: table() cross-tabulates the two factors, then chisq.test() does the rest. The high p-value here (0.91) means smoking patterns look similar across sexes in this sample.
How do you check the assumptions before trusting the result?
Three assumptions sit underneath every chi-square test of independence, and skipping the checks is the single most common mistake practitioners make. Let's walk them.
- Independent observations. Each count in the table comes from a separate, independent unit (one row per person, one survey response per person). Repeated measures or matched pairs break this assumption, you'd use McNemar's test instead.
- Expected counts large enough. Every cell's expected count should be at least 5. A common relaxation: at most 20% of cells may have expected counts below 5, and no cell should have expected count below 1.
- Fixed categories with random sampling. Categories are defined before data collection, and rows of the dataset are a random sample of the population.
The first and third are study-design questions. The second is something you must verify from R every single time.
Half the cells have expected counts below 5, and several cells (Heavy/None, Occas/None, Regul/None) have expected counts of 1 or below. This breaks the rule of thumb. Treat the p-value with suspicion until you re-run the test with a method that handles small expected counts (covered later in the Yates / simulate / Fisher section).
When the assumption is badly violated, R itself often warns you. Let's reproduce the warning on a clearly small table.
The warning Chi-squared approximation may be incorrect is R's way of telling you the asymptotic p-value cannot be trusted. It is not optional. Ignoring it can flip a significant result to non-significant or vice versa.
Try it: Write ex_check_expected(tbl) that returns TRUE if at least 80% of cells have expected count >= 5 AND no cell has expected count below 1. Test it on tbl from earlier and on small_tbl.
Click to reveal solution
Explanation: chisq.test(tbl)$expected returns the expected-count matrix without you having to recompute it. mean(e >= 5) is the share of cells meeting the floor, and all(e >= 1) enforces the absolute minimum. Both must hold.
What do the chi-square statistic, p-value, and degrees of freedom mean?
Once the assumptions hold, the three numbers in the output have specific roles:
- Chi-square statistic ($\chi^2$) measures how far observed counts sit from expected counts, summed across all cells. Bigger means more departure from independence.
- Degrees of freedom (df) is
(rows - 1) * (cols - 1). It captures the number of cells in the table that are free to vary once the row and column totals are fixed. - p-value is the probability of seeing a chi-square statistic at least as extreme as yours if the null (independence) were true.
The formula is small enough to read in one line:
$$\chi^2 = \sum_{i,j} \frac{(O_{ij} - E_{ij})^2}{E_{ij}}$$
Where:
- $O_{ij}$ = observed count in row $i$, column $j$
- $E_{ij}$ = expected count under independence: $\frac{\text{row}_i \text{ total} \times \text{col}_j \text{ total}}{\text{grand total}}$
- The sum runs over every cell in the table.
That's it, no fitting algorithm, no iteration. Pull the components from the fitted object to see the math directly.
Pearson residuals turn the global statistic into a per-cell story. A residual near 0 means that cell behaved as expected; a large positive residual means more observations landed there than expected; a large negative one means fewer. The biggest values flag where the action is.
For tighter tail probabilities, use standardized residuals. They have approximate variance 1, so values outside |2| are roughly the chi-square equivalent of a 5%-level z-score.
The largest standardized residual is in the Occas/Freq cell, but at 1.66 it does not exceed |2|, consistent with the non-significant overall test. If you ever see a standardized residual of 3 or 4 in a non-significant test, double-check, you may have an interaction worth investigating.
Try it: From xt, extract the row name and column name of the cell with the highest absolute standardized residual. Save them to ex_row and ex_col.
Click to reveal solution
Explanation: which(..., arr.ind = TRUE) returns row/column indices for matrix entries; you then look up the names from rownames() and colnames() of the residual matrix.
How big is the effect? Computing Cramer's V, phi, and contingency coefficient
The p-value answers "is the association real?". With 50,000 observations, even a tiny, practically meaningless association will return a p-value below 0.001. Effect size answers "is the association big enough to care about?". For r-by-c tables, the standard measure is Cramer's V.
$$V = \sqrt{\frac{\chi^2}{n \cdot \min(r-1, c-1)}}$$
Where:
- $\chi^2$ = the chi-square statistic from the test
- $n$ = total sample size (sum of all cells)
- $r$, $c$ = number of rows and columns in the table.
V ranges from 0 (no association) to 1 (perfect association). For 2x2 tables, V reduces to the phi coefficient, which is just $\sqrt{\chi^2 / n}$. The math is simple enough that you do not need a separate package, base R will do it.
Cramer's V of 0.10 is tiny. The chi-square test gave us p = 0.48 and now we know that even if it were significant, the practical association would be negligible.
The interpretation thresholds depend on the smaller dimension of the table. Cohen's commonly cited cutoffs are below.
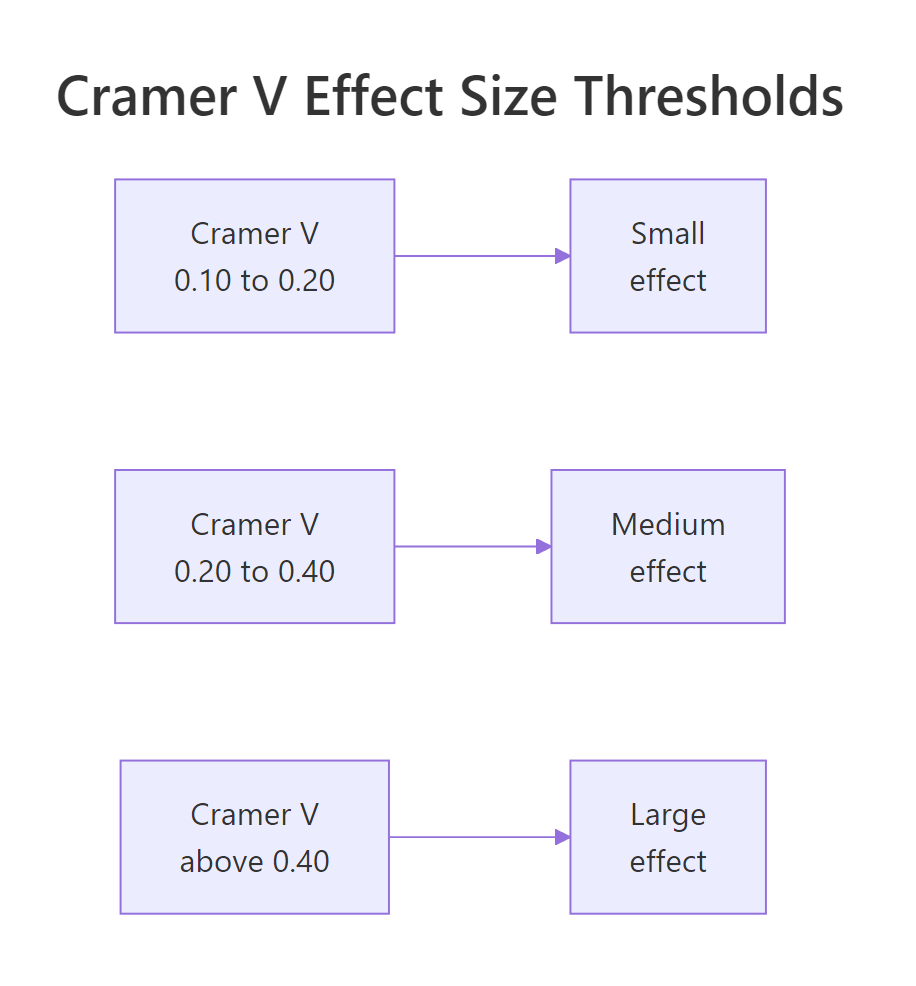

Figure 1: Cramer's V interpretation thresholds for small, medium, and large effects.
| df = min(r-1, c-1) | Small | Medium | Large |
|---|---|---|---|
| 1 (2x2 table) | 0.10 | 0.30 | 0.50 |
| 2 (e.g. 2x3, 3x3) | 0.07 | 0.21 | 0.35 |
| 3 (e.g. 2x4, 4x4) | 0.06 | 0.17 | 0.29 |
| 4 | 0.05 | 0.15 | 0.25 |
Use these as guidelines, not laws. A V of 0.12 is "small" by Cohen but might be the most important finding in your study, depending on context.
p < 0.001 and no V is reading half the result.Try it: Compute Cramer's V manually for ex_xt (the smoking-by-sex test you ran earlier). The total sample size is sum(ex_tbl). Save your answer to ex_v.
Click to reveal solution
Explanation: as.numeric() strips the named-vector wrapper from $statistic so the arithmetic stays clean. The min(r-1, c-1) term is what makes V comparable across tables of different shapes.
When should you use Yates correction, simulate.p.value, or switch to Fisher's exact?
R's default behavior changes with table size, and the choice of correction affects your p-value. Three knobs are worth knowing.
For 2x2 tables, chisq.test() applies the Yates continuity correction by default. This subtracts 0.5 from each |O - E| before squaring, which makes the test more conservative (bigger p-values). For larger tables, Yates does not apply.
For tables with sparse cells, you have two robust choices: Monte Carlo simulation of the p-value, or Fisher's exact test.
Yates pushes the p-value from 0.0028 to 0.0051, a meaningful shift near common decision thresholds. Statisticians have argued for decades over whether Yates is appropriate; modern practice is split. If you are unsure, reporting both is honest.
correct even if you set it. Many older tutorials do not mention this and produce confusing output for newcomers.When expected counts are too small, neither correction will fix the issue. Switch to a method that does not rely on the chi-square approximation at all.
Both give p-values around 0.78, very close to each other and well above 0.05. Either would be a defensible report; Fisher's is preferred for 2x2 tables and small overall samples (n < 20-30), while Monte Carlo scales better to larger sparse tables.
The decision tree below summarizes the choice.
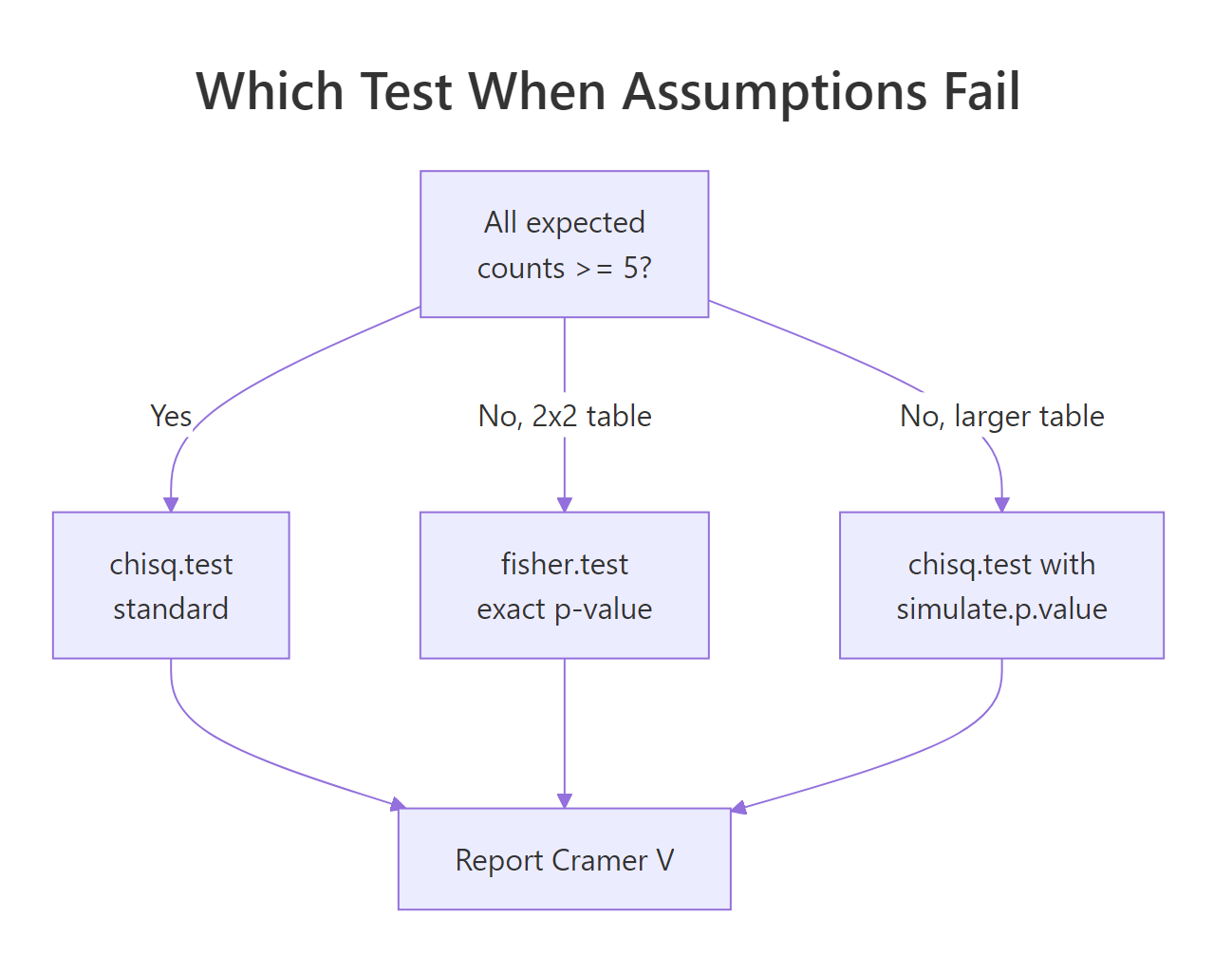
Figure 2: Decision tree for picking between standard chi-square, simulated p-value, and Fisher's exact.
Try it: You have a 2x2 table where one expected count is below 5. Run the appropriate test (hint: it is in the diagram above) and save the resulting p-value to ex_p.
Click to reveal solution
Explanation: With a 2x2 table and an expected count below 1, neither standard chi-square nor Yates fixes the problem. Fisher's exact uses the hypergeometric distribution to compute an exact p-value, no asymptotic approximation involved.
How do you compute statistical power and required sample size?
Power analysis answers two related questions:
- Post-hoc: Given my sample size, what effect size could I have detected with 80% power?
- A-priori: To detect an effect of a given size with 80% power, how big a sample do I need?
The chi-square test's power depends on the sample size $n$, degrees of freedom $df$, the alpha level, and the effect size $w$, which is a population analog of Cramer's V (small = 0.10, medium = 0.30, large = 0.50 for df = 1). The math uses the noncentral chi-square distribution. The popular helper for this is pwr::pwr.chisq.test(), but the underlying computation is two lines of base R via pchisq(), which is what we'll show here.
With n = 235 and df = 6, you have only 17% power to detect a small effect (w = 0.10). The non-significant result you saw earlier could just mean your study was underpowered, not that the effect doesn't exist. For a medium effect you have 95% power, that finding is trustworthy.
To turn the question around, find the smallest n that gives 80% power. Solve numerically with a small loop.
You need 107 observations to detect a medium effect (w = 0.30) on a df = 2 table with 80% power at alpha = 0.05. This is the kind of number you'd report in a grant application or pre-registration document.
Try it: Compute the sample size needed for w = 0.25, df = 4, alpha = 0.01, and power = 0.90. Save the answer to ex_n.
Click to reveal solution
Explanation: Stricter alpha (0.01 vs 0.05), higher target power (0.90 vs 0.80), and a smaller effect size (0.25 vs 0.30) all push n upward. 313 is what each of those constraints demands together.
Practice Exercises
These capstone exercises combine the steps above. Use distinct variable names (my_*) so they don't collide with the tutorial's notebook state.
Exercise 1: Sex and smoking, end-to-end
Using MASS::survey, test whether Sex and Smoke are independent. Report the chi-square statistic, df, and p-value, then compute Cramer's V. Save Cramer's V to my_v_sex_smoke.
Click to reveal solution
Explanation: A p-value of 0.91 plus V of 0.05 means there is essentially no relationship between sex and smoking in this dataset, a clear "no" on every dimension.
Exercise 2: Sparse 2x2, three different p-values
Build the 2x2 table below, where one expected count drops below 5. Run three tests: standard chisq.test() (Yates on), chisq.test(simulate.p.value = TRUE), and fisher.test(). Save the three p-values to my_p_chi, my_p_sim, and my_p_fisher. Which is most defensible to report?
Click to reveal solution
Explanation: The standard chi-square crosses 0.05 (looks "significant") but the warning fires. Simulated and Fisher both sit just above 0.05 and agree with each other. Fisher is most defensible here, exact, no approximation, and standard for sparse 2x2 tables.
Exercise 3: Sample size for a pilot
A pilot study estimated w = 0.25 (small-to-medium) on a contingency table with df = 4. You want 90% power at alpha = 0.01 for the main study. How many observations do you need? Save to my_n.
Click to reveal solution
Explanation: Same answer (313) as the inline exercise, the harder constraints on alpha and power matter more than the slightly larger df.
Complete Example
Here is the full pipeline on a different dataset, HairEyeColor, a 3-way array shipped with base R. We'll collapse over Sex, run the full test, and write a publishable summary.
Reportable summary (3 sentences):
A chi-square test of independence found a strong association between hair color and eye color, $\chi^2$(9, N = 592) = 138.3, p < .001. Cramer's V = 0.28, a medium-to-large effect. Standardized residuals show the largest deviations among Blond hair: substantially fewer brown-eyed (z = -10.5) and substantially more blue-eyed (z = 14.2) blondes than independence would predict.
That paragraph is the deliverable, statistic, df, n, p-value, effect size, and a sentence locating the association. It is what every reviewer wants to see.
Summary
The chi-square test of independence is a six-step pipeline, not a one-line call.
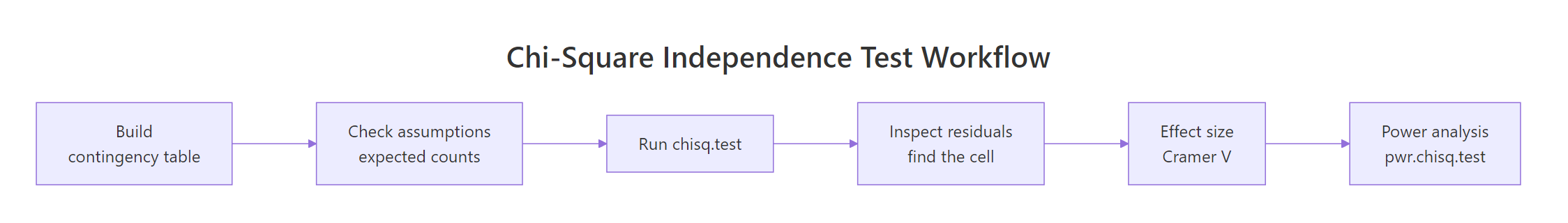
Figure 3: The full chi-square workflow, from contingency table to power analysis.
| Step | What to do | R function |
|---|---|---|
| 1. Build table | Cross-tab the two categorical variables | table() |
| 2. Check assumptions | Verify expected counts >= 5 in 80%+ of cells | chisq.test()$expected |
| 3. Run test | Compute $\chi^2$, df, p-value | chisq.test() |
| 4. Locate effect | Inspect standardized residuals | chisq.test()$stdres |
| 5. Effect size | Compute Cramer's V | $\sqrt{\chi^2 / (n \cdot \min(r-1, c-1))}$ |
| 6. Power | Post-hoc detection or a-priori sample size | pchisq() with ncp |
| Fallbacks | Sparse table | simulate.p.value or fisher.test() |
Three habits separate good practice from p-value chasing: always inspect expected counts, always report effect size, and run an a-priori power analysis before collecting data.
References
- R Core Team.
chisq.test: Pearson's Chi-squared Test for Count Data. R documentation. Link - Cohen, J. (1988). Statistical Power Analysis for the Behavioral Sciences, 2nd ed. Lawrence Erlbaum Associates.
- Agresti, A. (2018). An Introduction to Categorical Data Analysis, 3rd ed. Wiley.
- Champely, S. The
pwrpackage: Basic functions for power analysis. CRAN. Link - Yates, F. (1934). Contingency tables involving small numbers and the chi-square test. Journal of the Royal Statistical Society Supplement, 1(2), 217-235.
- Cramer, H. (1946). Mathematical Methods of Statistics. Princeton University Press.
- McHugh, M. L. (2013). The chi-square test of independence. Biochemia Medica, 23(2), 143-149. Link
Continue Learning
- Categorical Data in R: Frequency Tables, Crosstabs & Mosaic Plots, the foundation post on building contingency tables, mosaic plots, and reading proportions before you ever run a test.
- Chi-Square Goodness-of-Fit Test in R, the sister test, asking whether one categorical variable matches an expected distribution rather than asking whether two variables are related.
- Fisher's Exact Test in R, go deeper on the exact-test alternative recommended for sparse 2x2 tables.