R Data Types: Which Type Is Your Variable? (And Why It Matters)
R has six basic data types, numeric, integer, character, logical, complex, and raw, and the type of a value decides what you can do with it. Get it wrong and your math turns into string concatenation, your filters return nothing, or your model silently drops rows.
What are R's six data types?
Before you can trust a number, you need to know whether R actually sees it as a number. Every value in R belongs to exactly one of six basic types, and one call to class() tells you which. Let's create one value of each type and ask R to identify them.
Six values, six different types. weight is numeric because 72.5 has a decimal. age is integer only because we added the L suffix, without it, R would store 30 as numeric too. name is character (always quoted). is_raining is logical (the Boolean). signal is complex (has an imaginary part). byte is raw (low-level bytes). That's the whole family.
Try it: Create a variable ex_city holding the string "Chennai" and print its class.
Click to reveal solution
Explanation: Any value wrapped in quotes becomes a character, regardless of what's inside the quotes, even "123" is character, not numeric.
How do I check a variable's type?
Knowing the types exist is one thing; checking them in running code is another. R gives you two families of tools: class() returns the type as a string, and is.*() functions return TRUE or FALSE for a specific type. Use class() when you want to know which type a value is, and is.numeric() / is.character() / is.logical() when you want to test if it's a particular type inside an if statement.
The last line is the trap every beginner hits. "72.5" looks like a number but it's wrapped in quotes, so R stores it as character. is.numeric() correctly says FALSE. This is exactly why data read from CSVs with a single stray non-numeric cell comes in as character, more on that in the coercion section.
Try it: Check whether the comparison 5 > 3 produces a logical value.
Click to reveal solution
Explanation: Comparison operators (>, <, ==, !=) always return logical values, which is why they drop straight into if conditions and filter() calls.
What's the difference between class(), typeof(), and mode()?
R has three functions that all seem to answer "what type is this?" and they disagree just often enough to cause confusion. Here's the rule: class() tells you the object's high-level class (what it behaves like), typeof() tells you how R stores it internally, and mode() is a legacy base-R category that you'll rarely need. Seeing them side-by-side makes the difference obvious.
Look at the first two rows. 72.5 has class "numeric" but typeof "double", that's because in R, "numeric" is the user-facing name for the double-precision floating point type. And both 72.5 and 30L have mode "numeric" because mode lumps integers and doubles together. For day-to-day work, class() is the one you want; typeof() matters when you care about storage (integer overflow, memory); mode() is a holdover from S that you can mostly ignore.
class() tells you what an object acts like; typeof() tells you how R stores it. A data frame has class "data.frame" but typeof "list", because internally a data frame is a list of columns. Knowing this difference is how you debug "why doesn't my function work on this object?" bugs.Try it: Print typeof(5) and typeof(5L) side-by-side.
Click to reveal solution
Explanation: 5 with no suffix is stored as a double (R's default), while 5L with the L suffix forces integer storage. class() would also show the difference ("numeric" vs "integer").
Why does numeric really mean "double" in R?
When most languages say "number" they mean integer. R flips that: every number you type is a double (a 64-bit floating-point number) unless you explicitly ask for an integer. That's why class(5) returns "numeric" and typeof(5) returns "double". If you want an actual integer you must add the L suffix, 5L, or use as.integer(5). This matters more than it sounds, because doubles and integers behave differently at extremes.
The integer overflow is a real gotcha. If you've got a counter that might cross 2.1 billion, row counts, byte totals, millisecond timestamps, an integer column will silently become NA. Doubles go to roughly 1.8 × 10^308 before overflowing, so they're the safer default for anything that might grow. That's precisely why R picked double as the default: one fewer surprise for scientific users.
Try it: Create an integer variable ex_count holding 100 using the L suffix and print its class.
Click to reveal solution
Explanation: Without the L, 100 would be a double. The L tells R "this is a literal integer", the same idea as the L suffix for long in C or Java, where R's designers borrowed the notation.
How does R coerce types automatically?
R doesn't let values of different types sit side-by-side in the same vector, everything in c(...) must share one type. When you mix types, R silently coerces them all to the single most flexible type in the group, following a strict hierarchy: logical → integer → double → complex → character. Character always wins because any value can be written as text.
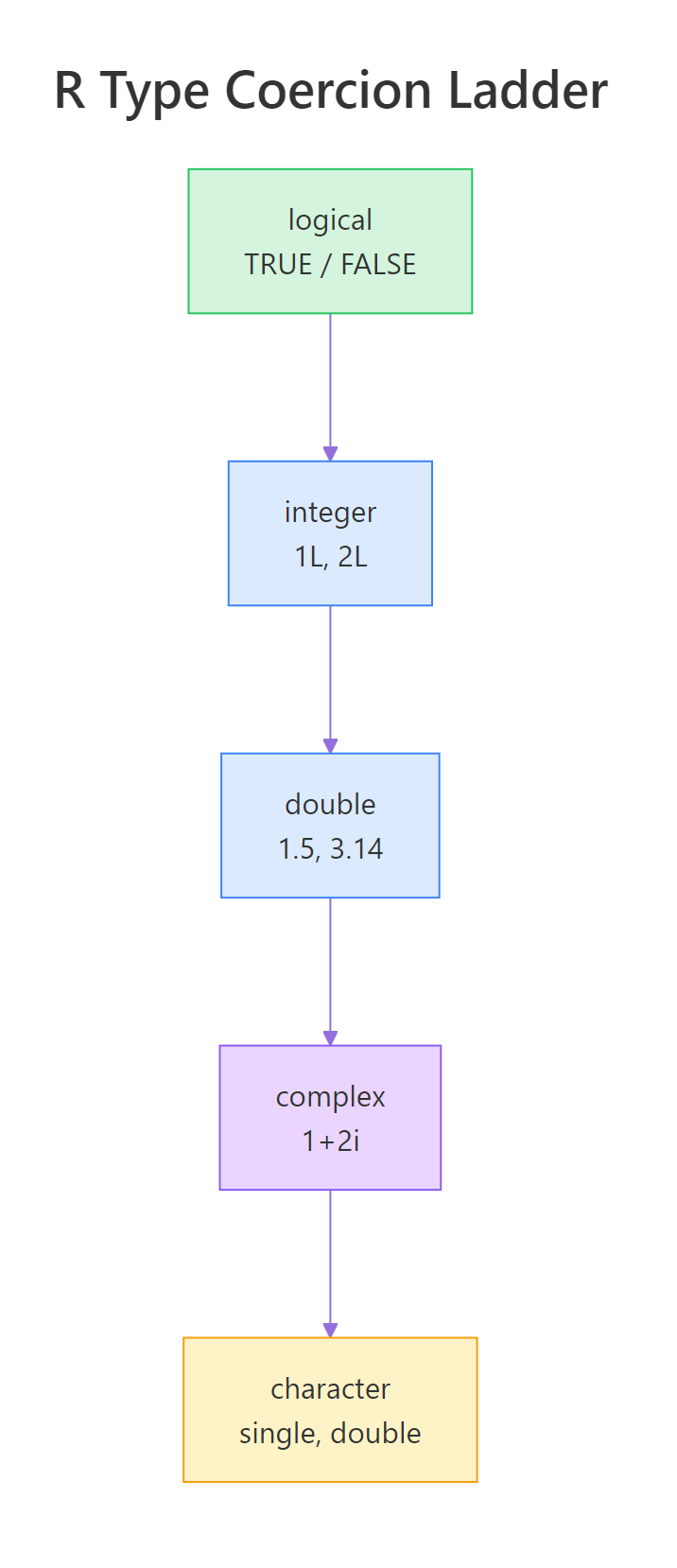
Figure 1: R's type coercion ladder, the "higher" type always wins when values are mixed in the same vector.
The ladder is easier to understand by example. Let's mix a number with a string and watch what happens.
That single "three" turned the other two numbers into strings "1" and "2", and sum() now throws an error. This is the single most common "my CSV won't add up" bug in R: one typo or one header row bleeding into the data makes the whole column character. The fix is either as.numeric(mixed) (which turns "three" into NA) or cleaning the source data.
Coercion also works the other way, when logicals meet arithmetic, R promotes TRUE to 1L and FALSE to 0L. That's how you count matches in a vector.
Three TRUEs means sum() returns 3, and mean() returns the proportion of TRUEs, a handy trick for computing "what fraction of my rows match this condition?" without a separate count.
class() your inputs first.Try it: Predict the class of c(TRUE, 1L, 2.5) before running the code.
Click to reveal solution
Explanation: Double is higher on the ladder than integer, which is higher than logical, so all three values coerce up to double. TRUE becomes 1.0 and 1L becomes 1.0.
What about NA, NULL, NaN, and Inf?
Not every value in R is a regular type, some represent "missing", "nothing", or "impossible." The four special values you'll meet are NA (missing), NULL (absent), NaN (not a number), and Inf (infinity). They look similar but behave very differently, and confusing them is a top source of bugs.
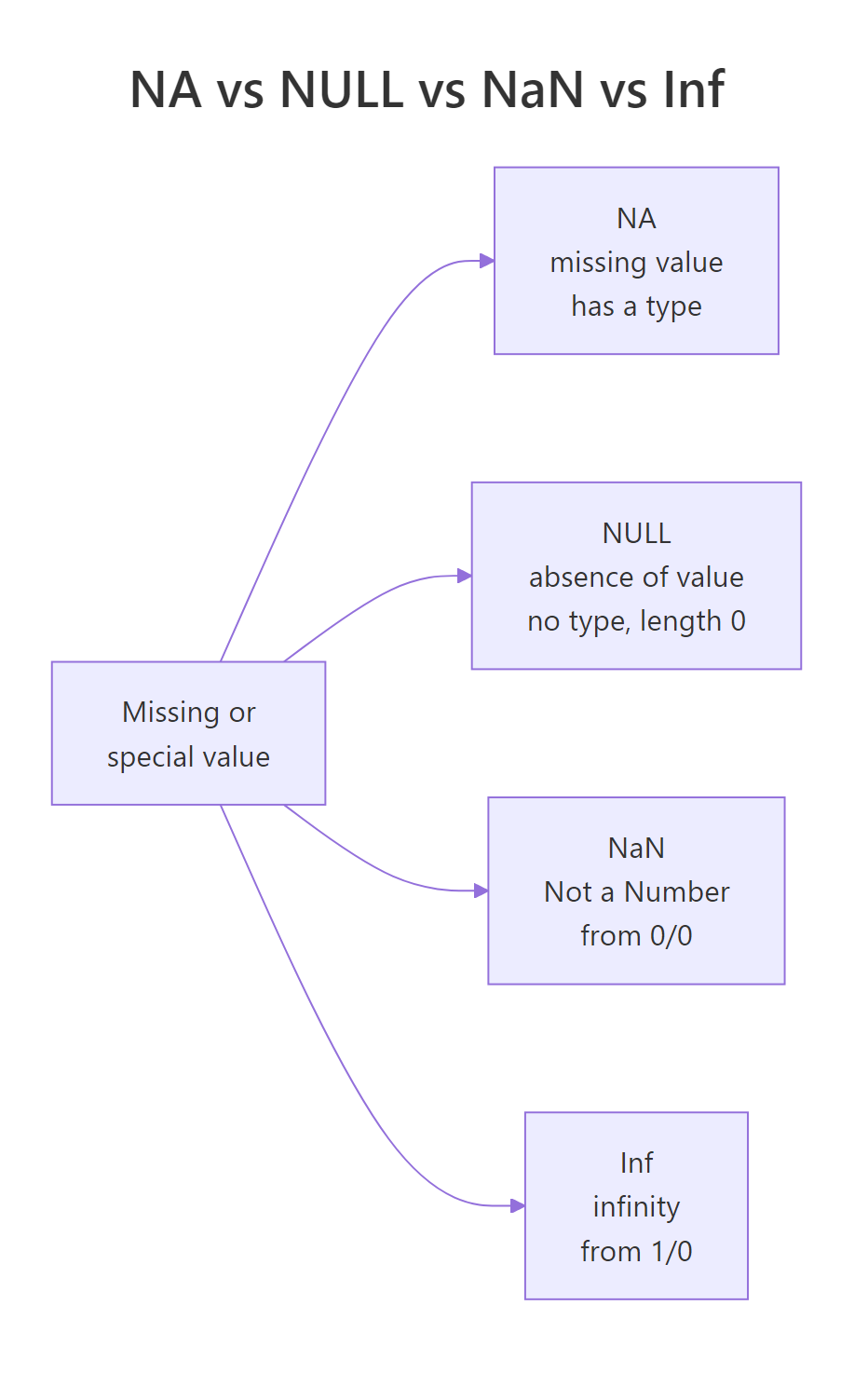
Figure 3: NA, NULL, NaN, and Inf, four different "missing-ish" values that behave very differently.
NA has length 1 and a type ("logical" by default, though typed variants like NA_integer_ exist). NULL has length 0, it's not a placeholder, it's truly nothing. NaN and Inf are both numeric; NaN comes from undefined operations like 0/0, Inf from divisions like 1/0. Your tests for them differ too: is.na() catches NA and NaN, is.null() catches only NULL, and is.infinite() catches Inf.
is.na(NULL) returns logical(0), not FALSE. Because NULL has zero length, is.na() returns a zero-length logical, which is neither TRUE nor FALSE and can crash if statements. Always use is.null() for NULL checks and is.na() for NA checks. Never mix them up.Try it: Count how many NAs are in the vector c(1, NA, 3, NA, 5).
Click to reveal solution
Explanation: is.na() returns a logical vector, and sum() coerces logicals to integers (TRUE → 1, FALSE → 0), giving you the count. This is the idiomatic R way to count missing values.
Which type should I use when?
With six types to choose from, the decision is usually obvious, but not always. Here's a practical decision tree: if the value is text, use character; if it's TRUE/FALSE, use logical; if it's a whole number that will never go near 2 billion, integer saves memory; otherwise use numeric (double). Complex and raw are specialist types, you'll know when you need them.
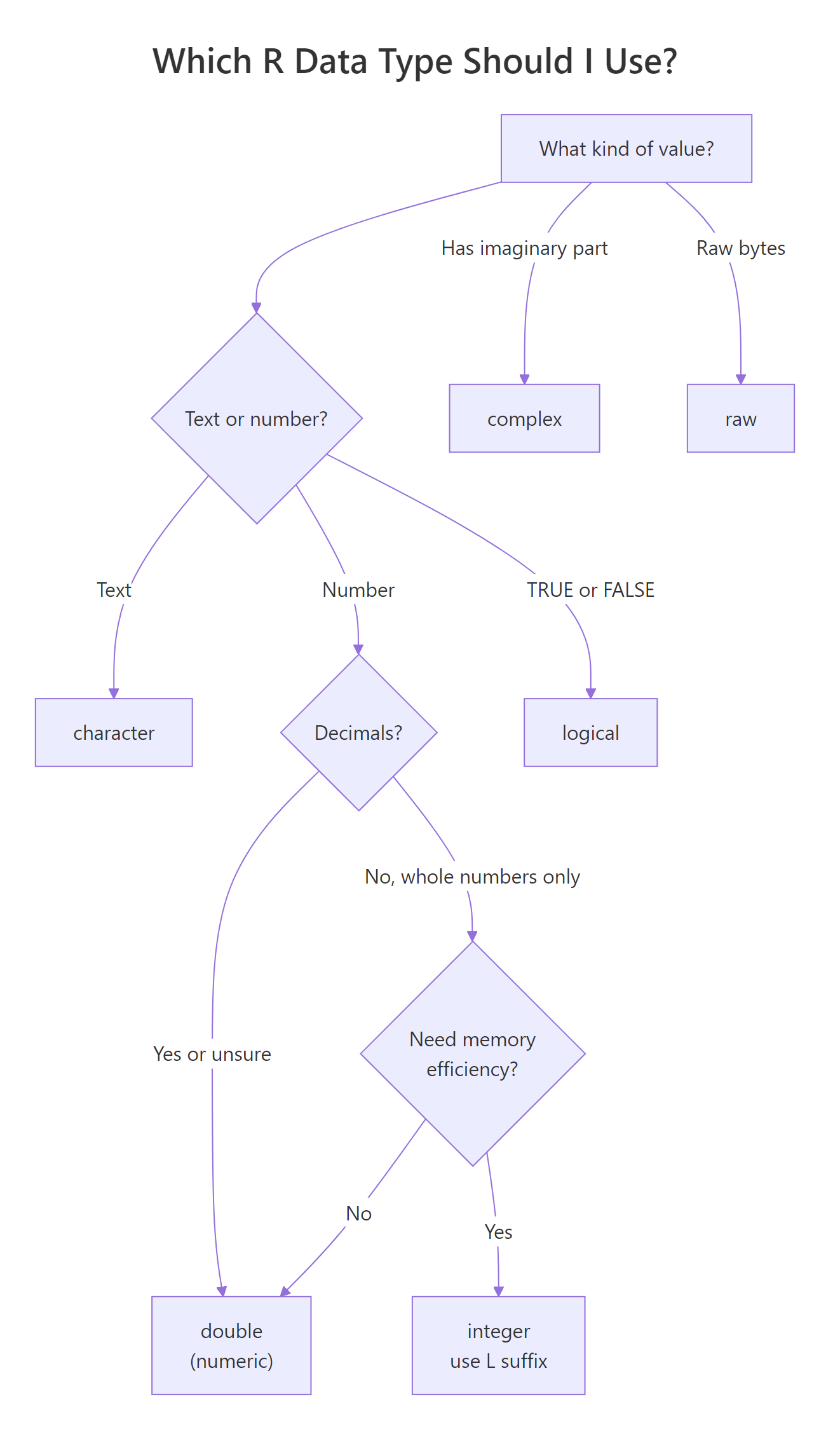
Figure 2: A decision tree for picking the right data type.
Ages are whole numbers, they never get huge, and you want memory efficiency when you've got millions of rows, so integer is the right pick. Notice that mean() still returns a decimal; R coerced the integer vector to double for the calculation. You get compact storage and accurate statistics.
Try it: Pick the right type for a vector of temperatures in Celsius and create it.
Click to reveal solution
Explanation: Temperatures can have decimals, so double (numeric) is the right pick. Using integer would force rounding and lose precision.
Practice Exercises
Time to put everything together. These capstones combine type checking, coercion, and picking the right type.
Exercise 1: Type detective
Given the vector my_values <- list(42, 42L, "42", TRUE, 4 + 0i), print the class() and typeof() of each element. Predict the answers before you run the code.
Click to reveal solution
Explanation: 42 and 42L look identical but have different storage types. "42" is character even though it looks like a number. 4 + 0i is complex even though the imaginary part is zero, the i forced the type.
Exercise 2: Fix the silent bug
The vector my_broken <- c(10, 20, "30") won't sum because the "30" poisoned the whole vector. Fix it by converting to numeric, then compute the sum as my_fixed.
Click to reveal solution
Explanation: as.numeric() coerces "30" back into a number. If the vector contained a genuine non-number like "three", you'd get NA with a warning, which is how R tells you the cast failed.
Exercise 3: Memory showdown
Create two vectors of 1 million values, one using integers (1L:1000000L) and one using doubles (1:1000000 * 1.0). Compare their memory footprints with object.size().
Click to reveal solution
Explanation: Integers use 4 bytes each; doubles use 8 bytes each. For a million elements that's a 4 MB difference, trivial on a laptop, but meaningful when you scale to hundreds of millions of rows.
Putting It All Together
Let's build a tiny weather log that uses four different types correctly, and then spot a contamination bug that beginners hit constantly.
Each column got the right type the first time: character for free text, numeric for measurements, logical for flags, integer for IDs. mean() and sum() work directly because the types match the operations. Then we broke temperature by inserting "--", a single bad cell made the whole column character, and mean() can't process strings. This is exactly what happens when you read messy CSVs, the fix is either readr::read_csv() (which reports parse errors) or as.numeric() with explicit NA handling.
Summary
| Type | Example | class() | typeof() | When to use |
|---|---|---|---|---|
| numeric | 72.5 |
"numeric" |
"double" |
Default for any number, measurements, statistics, money |
| integer | 30L |
"integer" |
"integer" |
Counts, IDs, indexes, saves memory at scale |
| character | "Selva" |
"character" |
"character" |
Any text, names, labels, categories |
| logical | TRUE |
"logical" |
"logical" |
Boolean flags, comparison results, filter masks |
| complex | 1 + 2i |
"complex" |
"complex" |
Signal processing, FFTs, rare in stats work |
| raw | as.raw(255) |
"raw" |
"raw" |
Binary files, bytes, cryptography, specialist use |
The three rules that catch 90% of type bugs:
- Check before you calculate.
class()andtypeof()are your friends, use them whenever something feels off. - One bad cell contaminates the whole vector. Mix a string in with numbers and R silently turns everything character.
- Doubles are the safe default; integers are an optimization. Only reach for
Lwhen you have millions of counts and memory matters.
References
- R Core Team, An Introduction to R, Section 2: Simple manipulations; numbers and vectors. Link
- Wickham, H., Advanced R, 2nd Edition, Chapter 3: Vectors. Link
- R documentation,
?numeric,?integer,?character,?logical. Run?typeofin R for full details. - Burns, P., The R Inferno, Circle 8: "Believing it does as intended" (coercion traps). Link
- Tidyverse,
readrtype guessing and thecol_typesargument. Link
Continue Learning
- R Vectors, how types combine with length to form R's most important data structure.
- R Type Coercion, a deeper dive on automatic conversions, gotchas, and the full coercion rulebook.
- R Special Values, full coverage of NA, NULL, NaN, and Inf with every function that handles them.