QR Decomposition in R: qr() & Why Regression Uses It Instead of Inverting
QR decomposition factors a matrix X into X = QR, where Q is orthogonal (its columns are perpendicular and unit length) and R is upper triangular. In R, you compute it with the base function qr(). Regression engines like lm() prefer QR over the textbook formula $\hat{\beta} = (X^\top X)^{-1} X^\top y$ because solving $R\hat{\beta} = Q^\top y$ by back-substitution is faster, and far more numerically stable, than inverting the cross-product $X^\top X$.
What does qr() actually return?
Many R guides mention QR factorization without ever opening the box. Let's create a small matrix X, call qr(), and pull out the orthogonal factor Q and the upper triangular factor R so we can see them on screen. Once you can read the output of qr(), every later trick (back-solving for regression coefficients, peeking inside lm()'s internals) becomes mechanical.
We start with a 3x3 matrix, factor it, and verify the product Q %*% R reconstructs the original matrix exactly.
Three things to notice. The matrix Q has columns that are mutually perpendicular and each has length one, so t(Q) %*% Q equals the identity matrix. The matrix R has zeros below the diagonal, which is what "upper triangular" means. And their product reproduces X to machine precision, confirming the factorization is exact, not approximate.
Try it: Build any 4x3 matrix ex_M, factor it with qr(), then check that t(Q) %*% Q equals the 3x3 identity. (Use round(..., 10) to suppress floating-point noise.)
Click to reveal solution
Explanation: t(Q) %*% Q returns the 3x3 identity because the three columns of Q are mutually perpendicular unit vectors. That is the defining property of an orthonormal matrix.
Why does regression use QR instead of inverting X'X?
OLS gives you a beautiful closed-form formula: $\hat{\beta} = (X^\top X)^{-1} X^\top y$. The trouble is, that formula multiplies the design matrix by its own transpose, which squares the condition number. The condition number is a single scalar that measures how much a matrix amplifies tiny errors. When you square it, you square the amplification, and an already-touchy regression problem becomes far worse.
The picture below contrasts the two routes from a design matrix to a coefficient estimate. Both end at $\hat{\beta}$, but the QR road avoids the costly detour through $X^\top X$.
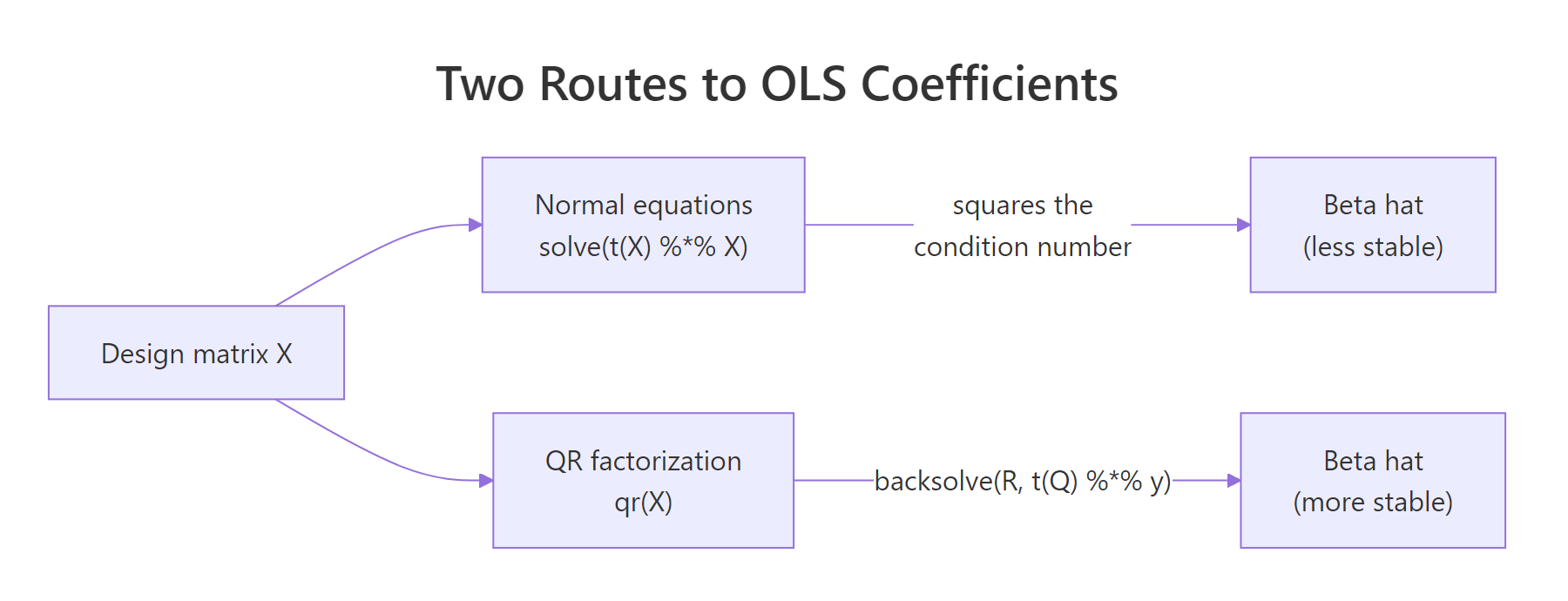
Figure 1: Two routes to OLS coefficients: normal equations vs QR factorization. The QR path avoids squaring the condition number.
Let's see the squaring effect on a near-collinear design matrix. Two of its columns are almost duplicates, which means $X$ is on the edge of being singular.
The condition number jumps from roughly $7 \times 10^5$ for X_design itself to about $5 \times 10^{11}$ for $X^\top X$. Notice the ratio is essentially the original condition number, confirming the squaring rule. Inverting $X^\top X$ for this problem amplifies floating-point errors by eleven orders of magnitude, which is enough to make several leading digits of the coefficients meaningless. QR works directly on X_design and never forms $X^\top X$, so the worst factor it ever inverts is R, whose condition number stays in the same ballpark as X itself.
Try it: Build your own ill-conditioned 3-column matrix ex_A (where one column is nearly the sum of the other two) and confirm that kappa(t(ex_A) %*% ex_A) is roughly kappa(ex_A)^2.
Click to reveal solution
Explanation: The ratio kappa(AtA) / kappa(A)^2 is close to one, which is exactly the squaring rule. The QR path stays at kappa(A), so it preserves about half the precision the normal equations would burn.
How do you solve least squares with QR?
Once you have X = QR, the least-squares problem $\min_\beta \lVert y - X\beta \rVert^2$ collapses to $R \hat{\beta} = Q^\top y$. The matrix R is upper triangular, so you can solve it with backsolve() instead of inverting anything. The whole thing fits in three lines.
We will reuse X_design from the previous block, generate a response y, run the QR pipeline, and check the result against lm().
The two columns agree to all printed digits. That is the practical proof that lm() and the manual QR pipeline solve the same equation, just through different code paths. The advantage of writing it out yourself is that you can see exactly which step would have failed if X_design were truly singular: the backsolve() call, where a zero on the diagonal of R would refuse to divide.
qr.coef() and qr.solve() skip the manual unpacking. If you do not need Q and R separately, qr.coef(qr_fit, y) returns the same beta in one call. For a one-shot solve from raw inputs, qr.solve(X_design, y) packages the entire pipeline.Try it: Using beta_qr, X_design, and y from above, compute the residual sum of squares manually as $\lVert y - X\hat{\beta} \rVert^2$ and store it in ex_rss.
Click to reveal solution
Explanation: Subtract the fitted values $X\hat{\beta}$ from y to get residuals, then square and sum. This is the quantity the OLS optimisation minimises, and it should match sum(residuals(lm(y ~ X_design - 1))^2).
How does lm() use QR under the hood?
When you call lm(y ~ x1 + x2), R does not invert anything. It computes a QR factorization of the design matrix and stashes it inside the fitted model object. You can reach in and inspect it.
The diagram below shows what qr() actually packs into its return value, and how the helpers (qr.Q, qr.R, qr.qty, backsolve) cooperate to produce $\hat{\beta}$.
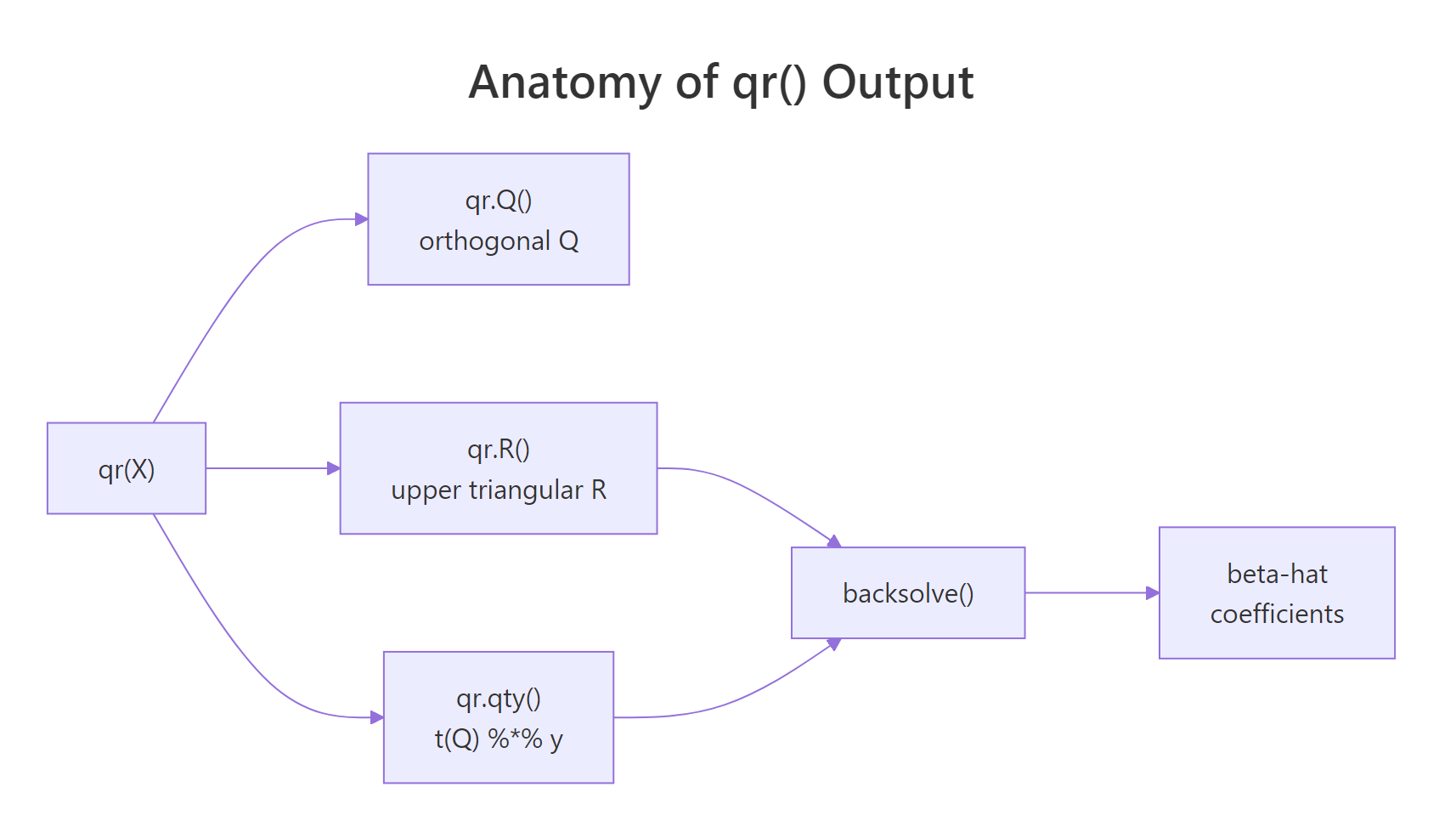
Figure 2: What qr() hands back: orthogonal Q, upper-triangular R, and helpers that combine them into beta-hat.
Now let's open up an lm() fit and confirm that its coefficients come straight out of the same QR primitives we just used by hand.
fit_lm$qr is the same qr object you would get from a direct qr(X_design) call (it carries qr, rank, qraux, pivot). The helpers qr.coef(), qr.fitted(), and qr.resid() re-derive the standard regression outputs from that single factorization. R never builds, never stores, and never inverts $X^\top X$.
fit$qr$qr stores Q and R packed into a single matrix. The output of qr() keeps the upper triangle of R and uses the strictly lower triangle to encode the Householder vectors that build Q. That is why qr.Q() and qr.R() exist as accessors instead of being plain matrix slots.Try it: Pull Q directly out of fit_lm (without calling qr() again) and check it has 3 columns and 100 rows.
Click to reveal solution
Explanation: qr.Q() accepts any object of class qr, including the one stored at fit_lm$qr. The result is the same orthonormal Q we extracted earlier, with 100 rows (one per observation) and 3 columns (one per predictor).
How is QR computed: Householder vs Gram-Schmidt?
There are two famous algorithms for building Q and R: classical Gram-Schmidt and Householder reflections. Gram-Schmidt is the textbook recipe (subtract projections, normalise) and is easy to write in a few lines of R. Householder is what real-world numerical libraries (LAPACK, LINPACK, base R's qr()) actually use because it is more stable in finite-precision arithmetic.
We will implement classical Gram-Schmidt by hand on a small 3x2 matrix, then compare to qr(). Reading the loop is the fastest way to internalise what "orthogonalize and normalise each column in turn" really means.
The two Q matrices differ only in the sign of each column, which is the non-uniqueness we flagged earlier. Both are valid QR factorizations of gs_X. The benefit of seeing Gram-Schmidt as code is that the loop makes the geometry obvious: each column of Q is the next column of X after subtracting away its overlap with the columns already accepted.
X are nearly parallel, floating-point cancellation in the subtraction step produces a Q whose columns are not quite orthogonal. Modified Gram-Schmidt and Householder reflections fix this. Real R code (the qr() function) uses Householder.Try it: Apply your own classical Gram-Schmidt to the matrix ex_GS_X below and verify the resulting ex_GS_Q has orthonormal columns.
Click to reveal solution
Explanation: The first column of ex_GS_Q is the first column of ex_GS_X rescaled to unit length. The second column subtracts the projection of ex_GS_X[,2] onto that direction before rescaling, so the two columns end up perpendicular and t(Q) %*% Q returns the 2x2 identity.
How do you handle rank-deficient or near-singular systems?
Real datasets contain perfect collinearities (a dummy variable equals the sum of others, a duplicate column slipped in, a continuous predictor times a constant). When that happens, X is rank deficient: one or more columns are linear combinations of the rest, and $X^\top X$ is singular. R's qr() is built for this case. It pivots columns by importance and reports the numerical rank in qr_obj$rank.
We will deliberately build a 4-column design where column 4 equals column 2 plus column 3 (a textbook collinearity) and look at how qr() flags it.
qr()$rank returns 3, not 4, because numerically one column adds no new direction. The pivot vector tells you which columns were kept first and which were demoted to the end: column 4 is last, which is the one R will treat as redundant. lm(y ~ X_rd - 1) returns NA for that coefficient for exactly this reason.
tol = to control how aggressive rank detection is. qr(X, tol = 1e-12) is stricter; qr(X, tol = 1e-3) is more lenient. The default of 1e-7 is calibrated for typical double-precision data. If you are working with very ill-conditioned predictors, lower the tolerance, but be ready to interpret near-collinearity warnings as a feature, not a bug.Try it: Build a 4x3 matrix ex_Xrd whose third column is twice its first column, then read off qr(ex_Xrd)$rank.
Click to reveal solution
Explanation: Even though the matrix has 3 columns, only 2 are linearly independent (column 3 is a scalar multiple of column 1). qr() returns rank 2, which is the same number of independent regressors lm() would actually fit.
Practice Exercises
These exercises combine multiple sections into harder tasks. Use distinct variable names (my_*) so they do not overwrite the tutorial's working state.
Exercise 1: Fit OLS on mtcars via QR and confirm against lm()
Using mtcars, build a design matrix my_X with columns (intercept, wt, hp, cyl) and a response my_y from mpg. Use qr(), qr.Q(), qr.R(), and backsolve() to compute beta_my. Compare against coef(lm(mpg ~ wt + hp + cyl, data = mtcars)). Both vectors should agree to machine precision.
Click to reveal solution
Explanation: model.matrix() adds the intercept column and lays out predictors exactly the way lm() would. The QR pipeline reproduces the same coefficients to all printed digits, confirming lm() is built on top of qr().
Exercise 2: Detect collinearity in a near-singular design
Construct a 5-column design matrix my_design (with 60 rows) where column 5 equals column 2 plus a tiny amount of noise: my_design[, 5] <- my_design[, 2] + 1e-9 * rnorm(60). Compute my_qr <- qr(my_design) and answer:
- What is
my_qr$rankwith the default tolerance? - Does it change if you set
tol = 1e-12? - Which column is dropped (last in
pivot)?
Click to reveal solution
Explanation: With the default tolerance of 1e-7, the near-duplicate column 5 is below the threshold for "new direction" and the rank is 4. Tightening the tolerance to 1e-12 lets the small numerical difference count, so the rank rises to 5. The pivot vector ends with 5, telling you that column was demoted as redundant. In practice, a rank that changes when you tweak tol is a strong sign of unstable predictors.
Complete Example
Here is the full QR-based regression pipeline on mtcars, with every output cross-checked against lm(). This block stitches together what we have been doing piece by piece.
The QR pipeline matches lm() on coefficients, fitted values, and residual sum of squares. You now have a transparent regression engine you wrote yourself in seven lines.
Summary
QR decomposition is the workhorse behind R's regression internals. The mindmap below recaps how the pieces fit together.
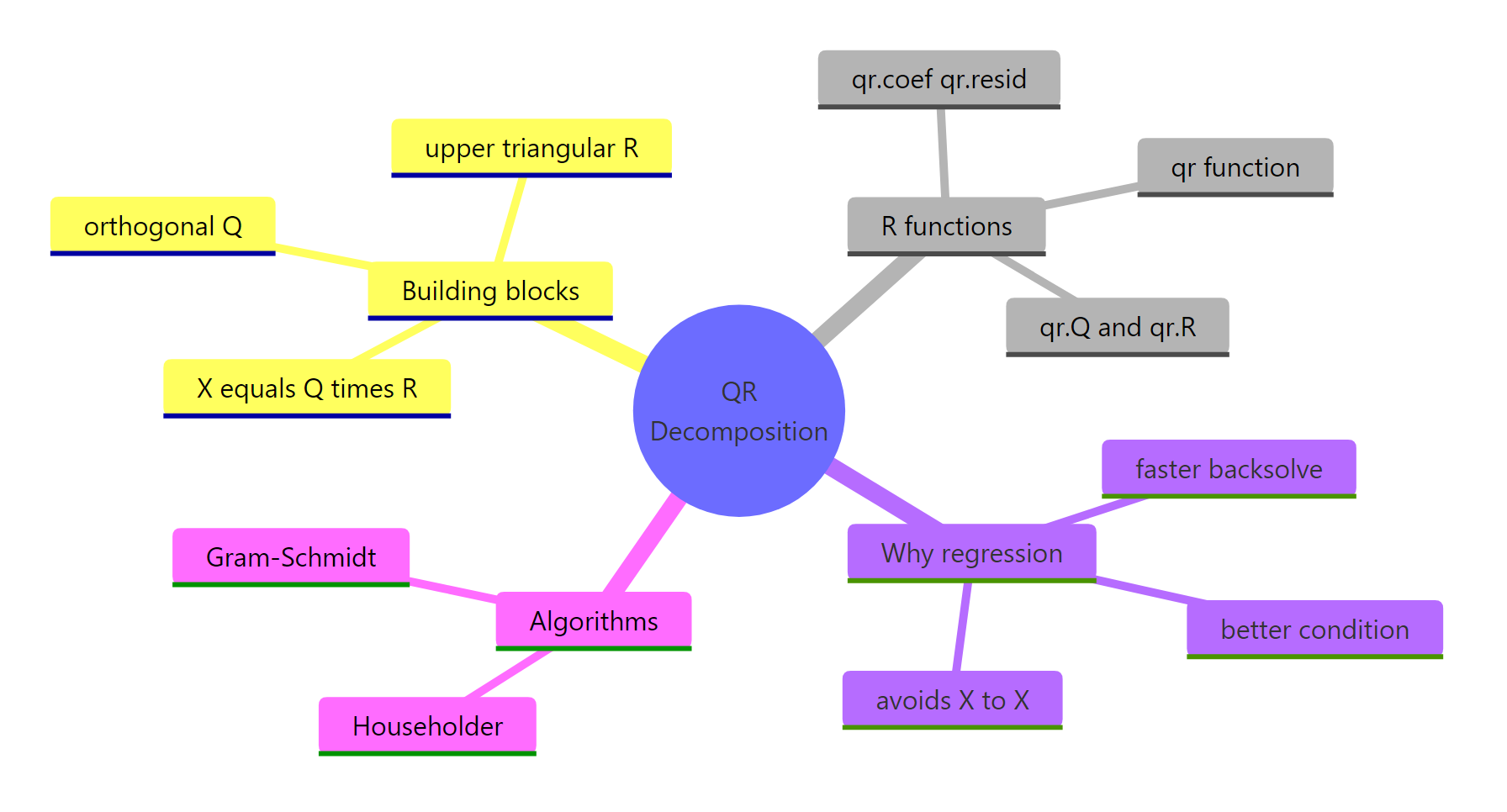
Figure 3: Big-picture map of QR decomposition concepts, R functions, and algorithms.
| Concept | What to remember | Function |
|---|---|---|
| Factorization | $X = QR$ with orthogonal $Q$ and upper triangular $R$ | qr(), qr.Q(), qr.R() |
| Why for regression | Avoids squaring $\kappa(X)$ that $X^\top X$ would cause | qr.coef(), qr.solve() |
| Inside lm() | fit$qr is a stored qr object; helpers reuse it |
qr.fitted(), qr.resid() |
| Algorithms | Householder is what base R uses; Gram-Schmidt is the textbook version | qr() (LINPACK / LAPACK) |
| Rank-deficient | qr()$rank and qr()$pivot reveal collinear columns |
tol = argument |
solve(t(X) %*% X) %*% t(X) %*% y, write qr.solve(X, y) instead. The QR call is a one-liner, runs no slower, and avoids the conditioning trap that has bitten OLS implementations for half a century.References
- R Core Team , qr: The QR Decomposition of a Matrix. R help page. Link
- Trefethen, L. N. & Bau, D. , Numerical Linear Algebra. SIAM (1997). Lectures 7-10 (Householder, QR, conditioning).
- Wickham, H. , Advanced R, 2nd Edition. CRC Press (2019). Link
- Larget, B. , The QR Decomposition and Regression, Statistics 496, UW-Madison. Link
- Irizarry, R. , Genomics Class: The QR decomposition. Link
- Strang, G. , Introduction to Linear Algebra, 5th Edition. Wellesley-Cambridge (2016). Chapter 4: Orthogonality.
Continue Learning
- Solving Linear Systems in R: solve(), qr() & Least Squares Solutions , when to call
solve()vsqr.solve()for square and overdetermined systems. - Projections & the Hat Matrix in R: OLS Geometry Explained Visually , how the hat matrix $H = X(X^\top X)^{-1} X^\top$ becomes a clean projection once you know QR.
- Singular Value Decomposition in R: svd() & The Most Useful Matrix Operation , SVD generalises QR and handles rank-deficiency without pivoting.