Aligned Rank Transform for Factorial Designs in R: ARTool Package
The Aligned Rank Transform (ART) is a nonparametric procedure that lets you run factorial ANOVA, including interaction effects, on ranked data when normality fails. The R package ARTool automates the alignment and ranking steps so you can fit two-way (and higher) designs with art() and read main effects, interactions, and post-hoc contrasts the way you would from a regular ANOVA.
What is the Aligned Rank Transform?
Suppose you ran a 2×4 experiment and the residuals are skewed. A classical two-way ANOVA is no longer trustworthy, and Kruskal-Wallis can only handle one factor at a time. ART solves both problems in one model. We'll start with the canonical example from Higgins (1990), where moisture and fertilizer were crossed in a peat-pot trial, and watch ART pick up the moisture × fertilizer interaction in eight lines.
Both main effects and the interaction come out significant on ranked data. The interaction p-value (4.6e-04) is the result a plain kruskal.test() could never give you, because Kruskal-Wallis collapses every cell into a single one-factor comparison. ART preserves the factorial structure and produces the same F-table layout you already know how to read.
Try it: Re-run the ART ANOVA on the subset of trays numbered 1 through 6 only. Save the table to ex_anova_sub.
Click to reveal solution
Explanation: Half the trays means roughly half the observations, so F values shrink. The structure of the table is unchanged because ART works on whatever rows you hand it.
How do you fit ART in R with art() and anova()?
The art() workflow is three steps every time: transform, verify, analyze. The transform step builds a rectangular table of aligned and ranked responses, one column per effect. The verify step checks that alignment actually zeroed out the other effects. The analyze step is anova(), which selects between lm, aov, and lmer based on your formula.
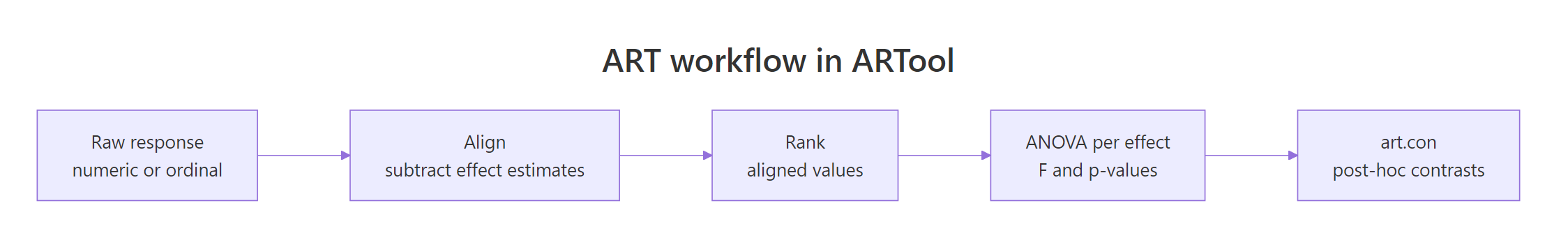
Figure 1: The three-step ART workflow (align, rank, ANOVA) followed by ART-C post-hoc contrasts.
Before any of that, the predictors must be factors. ART aligns by group means, so a numeric column will silently produce wrong results.
48 rows, 4 columns. Both predictors are already factors here, but if you load your own data with read.csv() you will likely need to coerce them.
as.factor() first and confirm with str().The four levels for each factor confirm we have a 4×4 design. Now fit the ART model and verify alignment with summary(). The verify step is the one most tutorials skip, but it is the only diagnostic ART gives you.
Three zeros where you want zeros. Column sums of zero confirm that aligning removed every other effect from each column, and F values of zero confirm that the residualized columns no longer carry signal from the other terms. If you ever see non-zero numbers here, the model is misspecified, usually because a predictor is numeric.
This is the same shape as aov() output: degrees of freedom, residual degrees of freedom, F, and a p-value. Read it the same way. The only thing to remember is that the response was the rank of an aligned column, not the raw DryMatter, so effect sizes need a partial-eta-squared formula rather than a raw mean difference.
Cohen's rule of thumb (small 0.01, medium 0.06, large 0.14) calls all three effects large. The interaction is the one that justifies using ART in the first place: a single-factor nonparametric test could not have produced a 0.56 partial eta-squared on the moisture × fertilizer term.
Try it: Coerce a copy of the dataset where Moisture is left as a character vector instead of a factor, then refit art(). Save the new ANOVA to ex_factor.
Click to reveal solution
Explanation: Recent ARTool versions auto-coerce character columns, but printing a warning. Always run as.factor() yourself so the model spec is explicit.
How do you run ART-C post-hoc contrasts?
Once the omnibus ANOVA is significant, you want to know which groups differ. The standard tool for this on a normal-theory ANOVA is emmeans::emmeans() followed by pairs(). On an ART model that is wrong: emmeans on the ART-fit gives marginal means of the aligned ranks, not the original response, and the pairwise tests are anti-conservative. ARTool ships its own contrast helper, art.con(), which implements the ART-C procedure of Elkin et al. (2021).
ART-C reports a t-ratio because it runs a regular t-test on the aligned-and-ranked column for each pair, then applies Tukey's HSD adjustment for the four-level family. M1 differs from M3 and M4 strongly; M3 versus M4 is not significant. The estimates are differences in mean ranks, so their absolute size depends on sample size, not the original units.
adjust = "tukey" for omnibus pairwise comparisons; switch to "none" only for planned contrasts. The default behaves correctly when you screen all pairs after a significant omnibus test. Setting adjust = "none" is appropriate only when you decided which contrasts to test before seeing the data.For interaction contrasts, name both factors separated by a colon. ART-C will compute the differences between every cell, which is usually too many comparisons unless you filter.
For 16 cells you get 120 unordered pairs, so the Tukey adjustment is severe. In practice, decide which interaction contrasts you actually care about (for example, the simple effect of Fertilizer within each Moisture level) and pass that subset to art.con() via the interaction = TRUE argument or filter the resulting data frame.
Try it: Run pairwise contrasts on Fertilizer (not Moisture). Save to ex_pair_fert.
Click to reveal solution
Explanation: Each main-effect family has $\binom{k}{2}$ pairs where $k$ is the number of levels; for 4 fertilizers that is 6.
How do you handle repeated measures and mixed effects?
Higgins 1990 was a split-plot: the same Tray received all four Fertilizer levels. Ignoring Tray treats those four rows as independent, which inflates degrees of freedom. ART supports both classical repeated-measures syntax (Error(Tray)) and modern mixed-effects syntax ((1|Tray)). The choice affects the underlying aov versus lmer fit, but the anova() interface is identical.
The mixed-effects route is the modern default because it handles unbalanced designs and missing cells without complaint.
Now Moisture is tested against 8 residual degrees of freedom (the between-tray scale) instead of 32, which is the correct denominator. The F drops from 23.85 to 4.56 because Moisture is being compared to tray-level variability, not the smaller within-tray noise. Fertilizer and the interaction sit on within-tray degrees of freedom (24 here), which are smaller than the original 32 because the random intercept absorbed some variance.
The classic split-plot syntax with Error(Tray) produces an aov fit. F values match the mixed-effects fit when the design is balanced.
Same numbers, different machinery. Use the mixed-effects form when the design is unbalanced or when you have multiple grouping variables; use Error() only if you specifically need an aov object for downstream tools that expect it.
Try it: Refit the mixed-effects ART using a random intercept for Moisture instead of Tray. Save to ex_mixed_alt.
Click to reveal solution
Explanation: The exercise illustrates that which variable becomes the random effect matters statistically, even if R will fit any syntactically valid formula.
When should you choose ART over Friedman or Kruskal-Wallis?
ART is the right tool only when you have two or more factors and you care about their interaction, or when you have a factorial design with random effects. For one-factor designs, the older nonparametric tests are simpler and just as powerful.
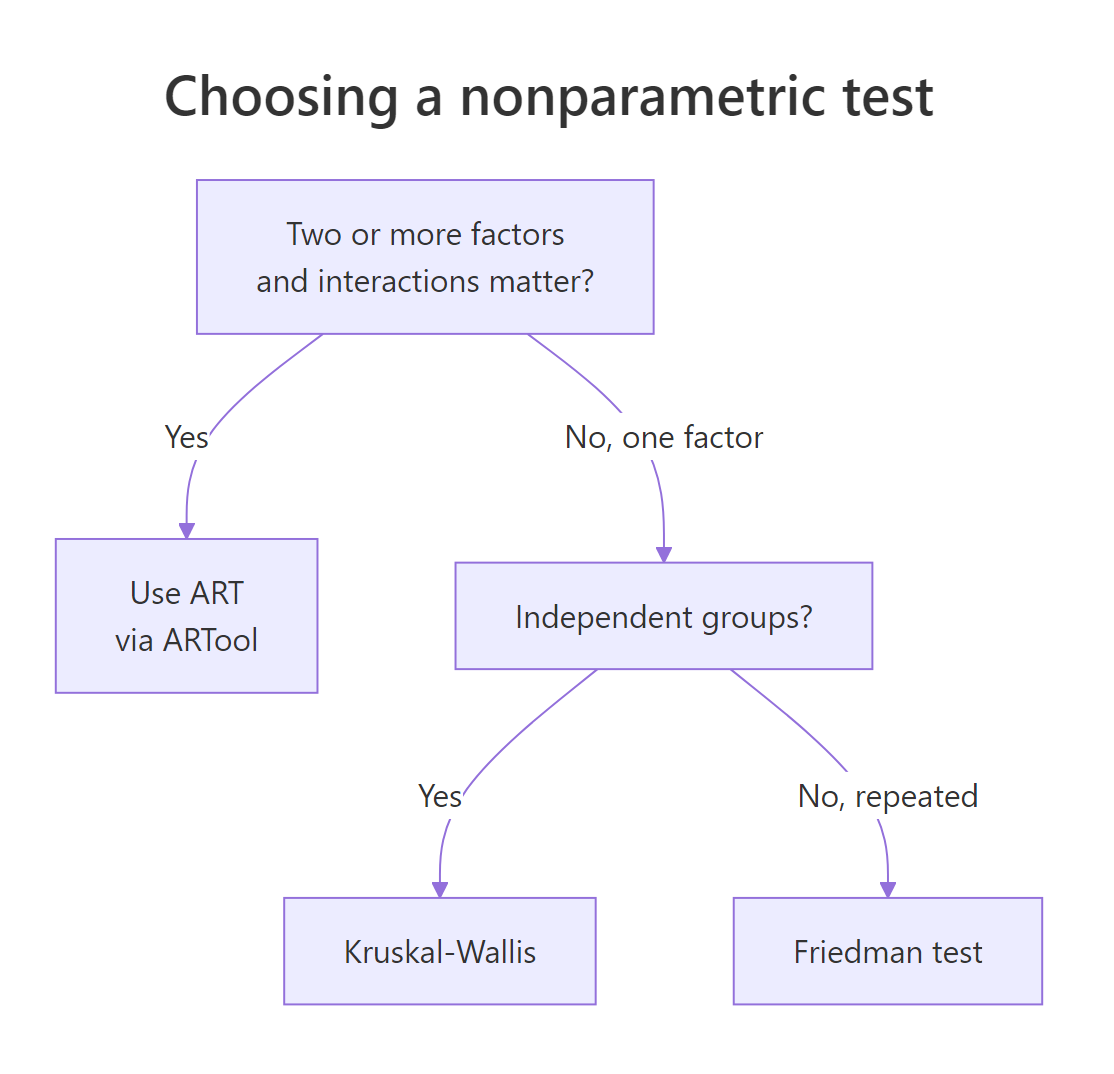
Figure 2: Choosing among ART, Kruskal-Wallis, and Friedman based on design.
The decision flow shrinks to three questions:
- Does my design have two or more factors and I want to test their interaction? Use ART via
ARTool. - Do I have one between-subject factor with three or more levels? Use Kruskal-Wallis (
kruskal.test). - Do I have one within-subject factor with three or more levels? Use Friedman (
friedman.test).
Classical parametric ANOVA is still preferable when residuals look approximately normal with constant variance. ART pays a small power cost for the rank transformation and is worth it only when those assumptions actually fail.
Try it: A study has one within-subject factor (Treatment) with three levels and 20 subjects, and you want to know if treatment differs. Pick the right test and assign it to ex_pick.
Click to reveal solution
Explanation: One within-subject factor with three or more levels is the textbook case for the Friedman test. ART would also work but adds machinery you do not need.
Practice Exercises
Exercise 1: ART on ToothGrowth
Run a full ART analysis on the built-in ToothGrowth dataset. The response is len, factors are supp (two levels) and dose (coerce to factor with three levels). Compute the F table, partial eta-squared for each effect, and Tukey-adjusted contrasts on dose. Save the ANOVA to tg_anova.
Click to reveal solution
Explanation: ART finds a significant interaction on ToothGrowth, which the rcompanion handbook reports too. Effect sizes label dose as a very large effect and the interaction as large.
Exercise 2: A rank-only interaction
Build a synthetic 3×3 dataset where the raw means show no interaction but the ranked responses do. The trick is to add a heavy-tailed noise component to one cell so its rank shifts dramatically while its mean barely moves. Fit ART, save the ANOVA to syn_anova, and confirm the interaction p-value is small.
Click to reveal solution
Explanation: The Cauchy noise produces a few extreme values that dominate ranks for cell (3, 3). ART picks this up on the ranked scale even though the cell mean is dragged toward those outliers and looks roughly continuous with the rest of the design.
Putting It All Together
The complete workflow in one block: load data, coerce factors, fit, verify, analyze, post-hoc. This is the script to keep in your snippets folder.
That is everything: a factorial nonparametric analysis with random effects and a defensible post-hoc procedure, in fewer than fifteen lines.
Summary
| What | How | |
|---|---|---|
| Run factorial nonparametric ANOVA | art(y ~ A * B, data = d) then anova(m) |
|
| Verify alignment | summary(m) returns column sums and F values that should be 0 |
|
| Test interactions | The A:B row in the ART ANOVA table |
|
| Mixed effects | Add `(1 | grouping) to the formula; ART routes through lmer` |
| Repeated measures | Add Error(grouping); ART routes through aov |
|
| Pairwise post-hoc | art.con(m, "Factor") with Tukey adjustment by default |
|
| Effect size | Partial eta-squared from F * Df / (F * Df + Df.res) |
References
- Wobbrock, J. O., Findlater, L., Gergle, D., & Higgins, J. J. (2011). The Aligned Rank Transform for Nonparametric Factorial Analyses Using Only ANOVA Procedures. CHI 2011. Link
- ARTool CRAN page. Link
- Kay, M. & Wobbrock, J. O. ARTool GitHub repository. Link
- Mangiafico, S. R Companion: Aligned Ranks Transformation ANOVA. Link
- Higgins, J. J., Blair, R. C., & Tashtoush, S. (1990). The aligned rank transform procedure. Proceedings of the Conference on Applied Statistics in Agriculture.
- Conover, W. J. & Iman, R. L. (1981). Rank transformations as a bridge between parametric and nonparametric statistics. The American Statistician, 35(3).
- Elkin, L. A., Kay, M., Higgins, J. J., & Wobbrock, J. O. (2021). An Aligned Rank Transform Procedure for Multifactor Contrast Tests. UIST 2021.
Continue Learning
- Kruskal-Wallis Test in R: the one-factor counterpart to ART, useful when interaction testing is not needed.
- When to Use Nonparametric Tests in R: decision rules for choosing between parametric and nonparametric procedures.
- Two-Way ANOVA in R: the parametric reference design that ART mimics on ranked data.