Aligned Rank Transform (ART) ANOVA in R: Factorial Nonparametric Tests
The Aligned Rank Transform (ART) is a nonparametric method that runs ordinary ANOVA on rank-transformed data, but only after aligning the data to isolate one effect at a time. It lets you test main effects and interactions in factorial designs when residuals violate normality, a gap that Kruskal-Wallis cannot fill. This tutorial uses the ARTool R package end-to-end.
When should you reach for ART instead of ordinary ANOVA?
Ordinary two-way ANOVA leans on normally distributed residuals. Real data, reaction times, Likert ratings, gene counts, rarely play along. Kruskal-Wallis rescues the one-factor case but has nothing to say about interactions. ART fills that gap: you keep the factorial structure, get main effects, an interaction F-test, and follow-up contrasts, all without assuming normality. Here is the one-minute version.
We will simulate a 2×3 factorial design where the response is strongly right-skewed (think reaction times in milliseconds). The formula has an interaction between Drug (A, B) and Dose (low, med, high). We fit art() and call anova() to read off the test statistics.
The output looks identical to a standard ANOVA table, but every F-value comes from a regression on ranks of aligned data. All three p-values are below 0.001: drug matters, dose matters, and the interaction is real. If you had tried aov() on this data, a Shapiro-Wilk test on the residuals would have rejected normality and invalidated the inference.
Try it: Generate a new 2×2 factorial with exponentially distributed response. Fit art() and confirm the interaction term appears in the ANOVA table.
Click to reveal solution
Explanation: Random data with no real effects produces non-significant F-values, exactly what you want as a sanity check. The three rows match the drug, dose, drug:dose structure from the main example.
What does "aligning" actually do to your data?
The word align sounds fluffy, but the math is simple: for each effect you want to test, subtract every other effect from each observation, then rank the residuals. The ranks you feed to ANOVA therefore reflect only the effect of interest.
Let's see this on a tiny 2×2 table. We have four observations, one per cell. To align for factor A, we need to strip out the effect of factor B and the AB interaction so that any variation left is due to A alone.
Notice what happened. The raw response bounced around (10, 12, 14, 20). The aligned response collapses to two values (12 and 17), one per level of A, exactly because alignment scrubbed out B and AB. The ranks then cleanly separate the two A-levels: 1.5 vs 3.5. ARTool performs this same trick automatically for every effect in the model.
Try it: Repeat the alignment for factor B using the formula aligned_B = y - cell_mean + B_marginal. Confirm the aligned values are identical within each B-level.
Click to reveal solution
Explanation: The two b1 observations now share one aligned value (11), and the two b2 observations share another (15). Alignment has isolated the B effect.
How do you run a full two-way ART ANOVA?
With the intuition in hand, the real workflow is three lines. We'll use Higgins1990Table5, a peat-pot experiment that ships with ARTool. It crosses four Moisture levels with four Fertilizer levels, with a Tray blocking variable.
Fit the model with art(), then always call summary() before interpreting results. The summary verifies that alignment worked: column sums of aligned responses should sit at 0, and F-values of ANOVAs on effects you are not currently testing should also be near 0. Non-zero values mean the model is misspecified.
Both diagnostics pass: the column sums are exact zeros and the "should-be-zero" F-values are all zero. Alignment is clean, so we can trust the anova() output next.
All three effects are significant. The interaction (F(9, 108) = 5.78, p < .001) tells you that the Fertilizer effect depends on the Moisture level; interpretation of main effects should therefore condition on that. This is the same reading rule as in parametric ANOVA.
anova() will still return an F-table, but its p-values will be wrong.Try it: Fit ART on the built-in ToothGrowth dataset using len ~ supp * dose. Cast dose to a factor first (it is numeric by default). Print anova().
Click to reveal solution
Explanation: Supplement, dose, and their interaction are all significant. Because the interaction is significant, the supplement gap depends on dose level.
How do you follow up with ART-C contrasts?
A significant F-test answers "is there some difference" but not "which groups differ." You'd normally reach for emmeans() + pairs() on a parametric ANOVA, but plain emmeans() on an ART model is biased: ranks were computed per effect, so estimated marginal means mix ranks that mean different things.
The fix is the ART-C procedure (Elkin et al., 2021). It re-aligns the data for each contrast before ranking, giving valid pairwise tests. In ARTool this is one call: art.con(). The function name's con is short for contrast.
Each row compares two cells of the Moisture×Fertilizer grid. Pairs with p.value < 0.05 indicate a genuine difference after Tukey adjustment. A cell like M1,F1 - M2,F1 (same Fertilizer, different Moisture) with p ≈ 1 tells you that switching Moisture level did not change DryMatter when Fertilizer = F1, which is the kind of nuance the interaction term hinted at.
art.con() for cells and art.con(m, "A") for main-effect contrasts, or artlm.con() when you need a custom emmeans object.Try it: Run art.con() on the main effect of Moisture using adjust = "bonferroni" instead of Tukey. How many of the six pairwise tests are still significant?
Click to reveal solution
Explanation: Four of the six pairs are significant at p < 0.05. Bonferroni is more conservative than Tukey, so you generally see fewer significant contrasts.
Does ART handle repeated measures and mixed models?
ART adapts to three design types through one formula interface. The presence or absence of an Error() term or a (1|Subject) term tells art() which underlying engine to use.
- Between-subjects factorial:
art(y ~ A * B, data = ...)fits vialm. - Repeated measures:
art(y ~ A * B + Error(Subject), data = ...)fits viaaov. - Mixed effects:
art(y ~ A * B + (1|Subject), data = ...)fits vialmerfrom thelme4package.
Here is a repeated-measures example where 30 subjects each rate a stimulus under every combination of Time and Condition.
The table reads the same way as before: main effects of time and condition plus a significant interaction, all derived from aligned-rank ANOVAs with subject as the error stratum.
art(y ~ A * B + (1|Subject)) works only if the lme4 package is installed. If your environment cannot load lme4, run the analysis in local RStudio after install.packages("lme4"). The Error()-based form shown above has no such dependency.Try it: Add a second within-subject factor session (two levels) to the repeated-measures formula and re-fit.
Click to reveal solution
Explanation: With no injected effect, all three p-values hover near 0.5, as expected for random exponential data. The formula pattern generalises to any number of crossed factors.
When should you NOT use ART?
ART is powerful, but it is not a free pass. Four situations argue against it.
- Cell size below ~10. Ranks carry little information in tiny cells. Consider bootstrap ANOVA (package
boot) or a GLM with a matching distribution instead. - Pure ordinal response with many ties. Proportional-odds models (
MASS::polr,ordinal::clm) respect the ordinal scale directly. - Residuals are already normal. Don't trade statistical power for nothing. Fit
aov(), checkshapiro.test()on the residuals, and only switch to ART if normality fails. - The "illusory promise" caveat. Earlier ART implementations produced poor contrast behaviour in some settings (see Elkin et al., 2021). The modern ART-C procedure (
art.con()) fixes this, but only if you use it instead of plainemmeans().
The quickest diagnostic is a side-by-side comparison: fit both models, check residuals of the parametric one, and confirm the ART p-values agree when assumptions hold.
Shapiro-Wilk rejects normality (p ≈ 10⁻¹⁸), so the parametric aov() is untrustworthy. Both analyses happen to agree on direction, but ART's p-values are the ones you'd report. If the Shapiro test had passed instead, the parametric ANOVA would be the better choice; there is no reward for using a rank test on data that cooperates.
aov(), plot residuals(aov0) or use shapiro.test(). If normality holds, stay parametric. If it fails, switch to art() and run the full workflow.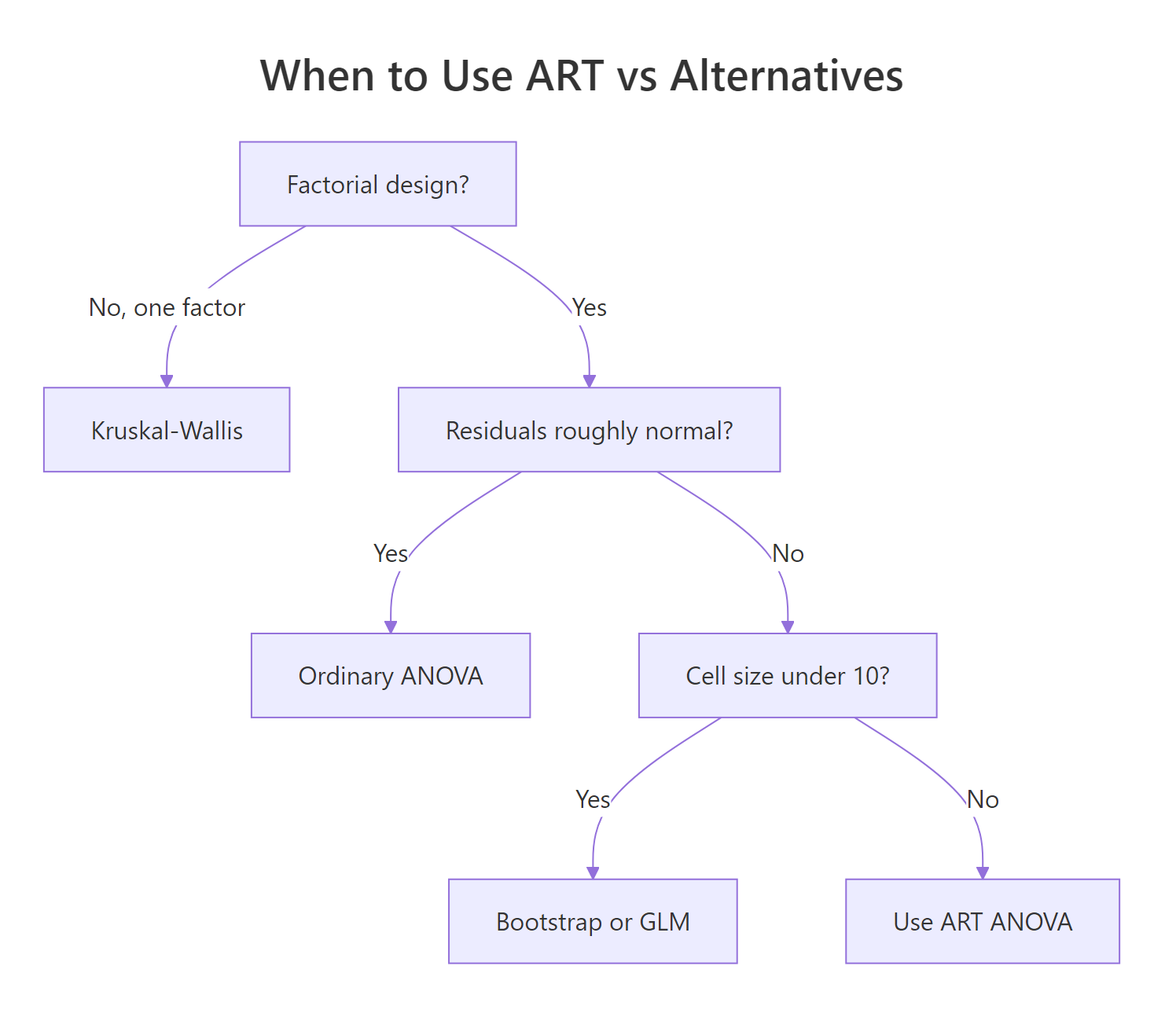
Figure 2: Decision guide for choosing between ART, ordinary ANOVA, Kruskal-Wallis, and GLM alternatives.
Try it: Simulate a 2×2 factorial with normal residuals, fit both aov() and art(), and confirm the p-values are close (within about 0.05).
Click to reveal solution
Explanation: The two interaction p-values are within 0.01 of each other, which is the expected behaviour when the parametric assumptions hold. ART does not lose power on normal data; it just doesn't add any.
Practice Exercises
Exercise 1: CO2 uptake factorial
Using the built-in CO2 dataset, fit an ordinary aov(uptake ~ Type * Treatment). Run shapiro.test() on its residuals. Then fit art(uptake ~ Type * Treatment) and print both ANOVA tables. Save the three p-values from the ART table to my_art_p.
Click to reveal solution
Explanation: Shapiro rejects normality (p ≈ 0.003), so ART is preferred. Type and Treatment are each significant, but the interaction is not (p ≈ 0.21).
Exercise 2: ToothGrowth contrasts
On ToothGrowth (cast dose to a factor), fit art(len ~ supp * dose), verify with summary(), and run art.con() on the interaction. Save the contrasts data frame to my_contrasts. Identify which supp-dose cells differ at α = 0.05.
Click to reveal solution
Explanation: art.con() returns the full pairwise grid; filtering on p.value < 0.05 extracts the significant cell differences after Tukey adjustment.
Exercise 3: Side-by-side comparison helper
Write a function compare_p(formula, data) that fits both aov() and art() with the given formula, pulls the p-values for each term, and returns a tibble with columns effect, p_parametric, p_ART. Test it on ToothGrowth with len ~ supp * dose.
Click to reveal solution
Explanation: On normally distributed data the two columns closely track each other. Wider gaps flag cases where the normality assumption is doing real work.
Putting It All Together
Here is the full ART workflow on ToothGrowth, from data check to interpretation.
Walking through the output: Shapiro is comfortably above 0.05 so the data happen to be roughly normal (ART agrees with parametric here, as expected). The ANOVA confirms significant effects of supp, dose, and the interaction. The pairwise contrasts pinpoint where the supp-dose effect lives: OJ at 0.5 mg/day is clearly lower than OJ at 1 or 2 mg/day, but at 0.5 mg/day OJ and VC don't differ. This is the kind of granular, defensible conclusion ART is built to produce.
Summary
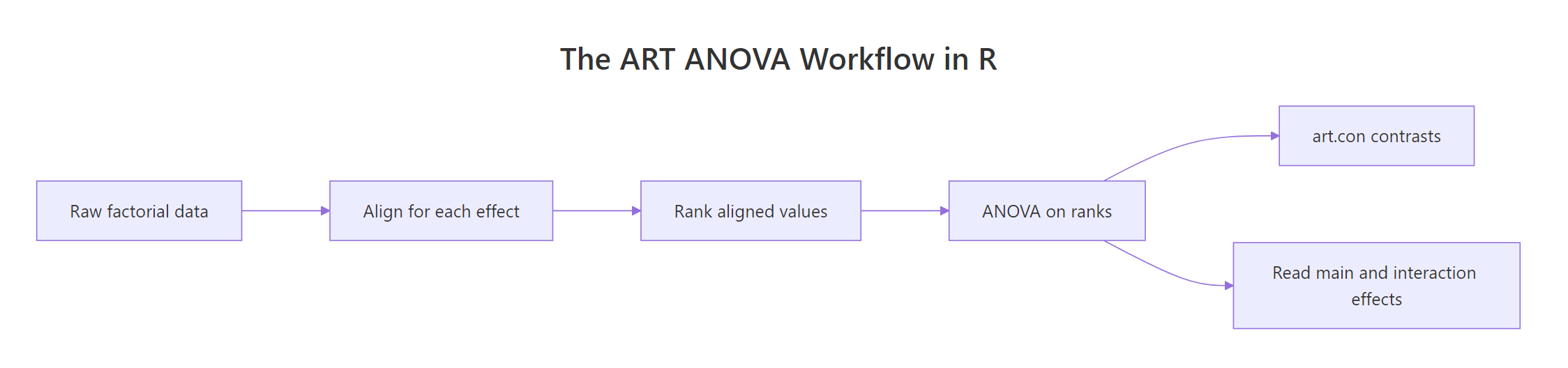
Figure 1: The ART ANOVA pipeline from raw data to pairwise contrasts.
| Step | Function | What it gives you |
|---|---|---|
| Fit | art() |
ART model object (lm, aov, or lmer engine) |
| Verify | summary(m) |
Column sums ~0 and aligned-F ~0 confirming clean alignment |
| Test | anova(m) |
F-statistics for every main effect and interaction |
| Contrast | art.con(m, "A:B") |
Pairwise post-hoc tests via the ART-C procedure |
Reach for ART when residuals fail normality and you need factorial structure (main effects plus interaction). Stay with parametric aov() when residuals are well-behaved. Use Kruskal-Wallis only for one-factor designs. Use art.con(), never plain emmeans(), for post-hoc tests on ART models.
References
- Wobbrock, J. O., Findlater, L., Gergle, D., & Higgins, J. J. (2011). The Aligned Rank Transform for Nonparametric Factorial Analyses Using Only ANOVA Procedures. Proc. CHI '11. Link
- Kay, M., Elkin, L. A., Higgins, J. J., & Wobbrock, J. O. (2025). ARTool: Aligned Rank Transform for Nonparametric Factorial ANOVAs. R package version 0.11.2. CRAN
- Elkin, L. A., Kay, M., Higgins, J. J., & Wobbrock, J. O. (2021). An Aligned Rank Transform Procedure for Multifactor Contrast Tests (ART-C). Proc. UIST '21. Link
- Mangiafico, S. S. (2016). R Handbook: Aligned Ranks Transformation ANOVA. rcompanion.org/handbook/F_16.html
- ARTool package vignettes:
vignette("art-contrasts"). GitHub repository - Higgins, J. J., & Tashtoush, S. (1994). An aligned rank transform test for interaction. Nonlinear World, 1(2), 201–211.
- ARTool reference manual (CRAN PDF). Link
Continue Learning
- Factorial Experiments in R: the parametric parent, 2^k designs with main effects, confounding, and interactions.
- Two-Way ANOVA in R: the classical factorial ANOVA that ART replaces when residuals break normality.
- Kruskal-Wallis Test in R: the one-factor nonparametric test that ART generalises to factorial designs.