Permutation ANOVA (PERMANOVA) in R: vegan Package for Ecology
PERMANOVA (Permutational Multivariate Analysis of Variance) tests whether the centroids of two or more groups differ in multivariate distance space. It uses permutations instead of normal-theory F distributions, so it works on species-abundance matrices, microbiome OTU tables, and any other multivariate data where ordinary MANOVA fails. In R, the vegan package implements it through adonis2().
What is PERMANOVA and when should you use it?
When you hold a multivariate response, like community species counts, microbiome OTU tables, or multi-trait measurements, and want to ask "do these groups differ overall?", PERMANOVA is the workhorse. It compares group centroids in distance space, builds the null distribution by shuffling group labels, and returns a pseudo-F and p-value. The classic dune dataset (20 sites, 30 species, four management types) is the standard worked example, and the call is one line.
The four management types explain about 34% of the multivariate variance in species composition (R² = 0.342). The pseudo-F of 2.77 is unusual enough that only 5 of 1000 permuted label-shuffles produced a value that large or larger, giving a p-value of 0.005. That is strong evidence that management regime drives community composition, not chance.
Try it: Refit the same model but use Moisture as the predictor instead of Management. Does soil moisture explain a similar share of community variance?
Click to reveal solution
Explanation: Moisture explains 40% of community variation, slightly more than Management. Both are real ecological drivers in this dataset.
How does PERMANOVA test differences using permutations?
Classical MANOVA assumes multivariate normality and equal covariance matrices. Real ecological data violate both. PERMANOVA replaces those assumptions with a single, simple idea: if your grouping does not matter, you should be able to scramble group labels and still get the same statistic on average. The pseudo-F is the same ratio you would compute in a one-way ANOVA, but applied to sums of squared distances rather than squared deviations.
If you are not interested in the math, skip to the next section, the practical takeaway is that p-values come from counting how often shuffled labels beat the observed F.
$$F = \frac{SS_B / (a - 1)}{SS_W / (N - a)}$$
Where:
- $SS_B$ = sum of squared distances between group centroids and the grand centroid
- $SS_W$ = sum of squared distances from each point to its own group centroid
- $a$ = number of groups
- $N$ = total number of observations
The block below pulls the permutation distribution out of adonis2() so you can see what Pr(>F) actually counts.
The histogram shows the null world. Almost every shuffled F lands below 2, but the observed F sits at 2.77, far in the right tail. The recomputed p-value of 0.006 matches the printed Pr(>F) from adonis2() (small differences come from the +1 continuity correction in vegan's count).
Try it: Refit the model with only 99 permutations, then with 9999, and compare the p-values. How much does the answer move?
Click to reveal solution
Explanation: With 99 permutations the granularity is 0.01, so the p-value cannot be smaller than that. 9999 permutations resolves the answer to four decimal places and removes most simulation noise.
How do you choose the right distance metric for PERMANOVA?
PERMANOVA is a test on a distance matrix, not on the raw data, so the metric you pick is part of the hypothesis. Three choices cover most cases: Bray-Curtis for species abundance counts (the ecology default, ignores joint absences), Jaccard for presence-absence data (set overlap), and Euclidean for continuous traits or transformed compositions. The diagram below maps data type to metric.
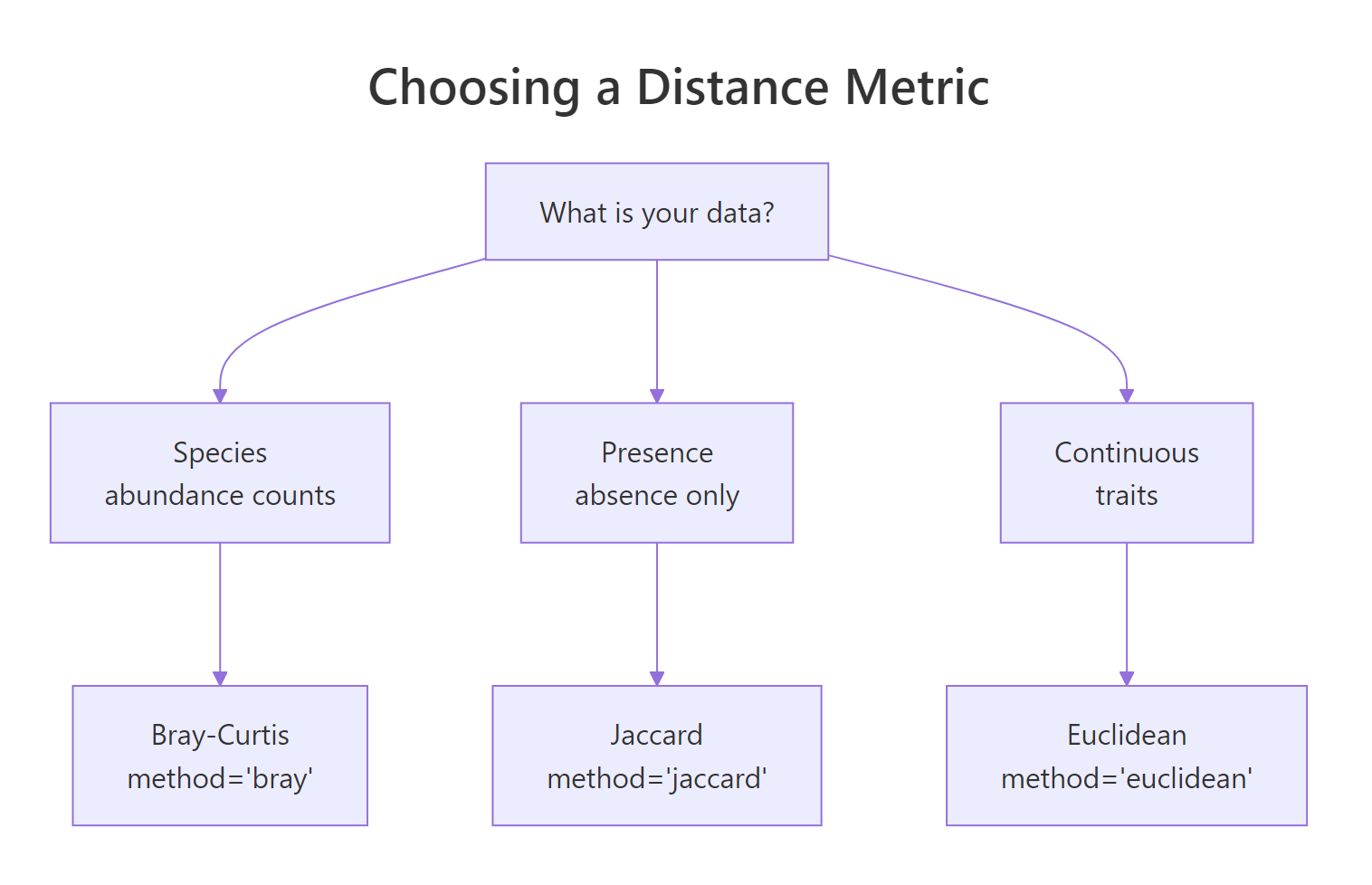
Figure 1: Choosing a distance metric for your data type.
Computing all three lets you see how the conclusion can shift with the metric.
All three metrics agree that Management matters, but the effect sizes differ. Bray-Curtis gives the largest R² because it weights the dominant species. Jaccard collapses abundance to presence-absence, so it loses some signal but stays significant. Euclidean treats raw counts as coordinates and is the least sensitive on this composition data, with a noticeably weaker p-value.
Try it: Run adonis2() directly on a Jaccard distance with binary = TRUE baked in via the formula interface, no precomputed vegdist.
Click to reveal solution
Explanation: adonis2() accepts method and binary arguments and computes the distance internally, so you do not need to call vegdist() first.
How do you check the homogeneity of dispersion assumption?
PERMANOVA's pseudo-F pools within-group sums of squares, which means it assumes the groups have similar multivariate spread. If one group is much more variable than the others, a significant adonis2 might be picking up a dispersion difference rather than a centroid difference. The vegan function betadisper() runs PERMDISP, the multivariate Levene test, on the same distance matrix.
The four management groups have between-group dispersions ranging from 0.23 (HF) to 0.43 (NM). The PERMDISP F is 2.0 with p = 0.15, well above 0.05, so we cannot reject the null of equal dispersion. Combined with the significant adonis2 result, this is the cleanest possible interpretation: groups differ in centroid location, not just in spread.
Try it: Run betadisper on Moisture instead and report whether dispersions are homogeneous.
Click to reveal solution
Explanation: Moisture groups are even more homogeneous in dispersion (p = 0.41), so the strong adonis2 effect on Moisture is purely a centroid-location story.
How do you run pairwise PERMANOVA and visualize results?
A significant adonis2 only tells you that some of the four management types differ. Pairwise PERMANOVA answers which pairs. The vegan package does not ship a built-in pairwise function, so the standard pattern is a small loop with combn() and a Bonferroni or BH correction.
Before correction every pair is nominally significant (p < 0.05), but six tests on the same data inflate the family-wise error rate. After Bonferroni correction no pair survives the 0.05 cutoff, although BF vs NM and NM vs SF sit just above. The honest read is that the overall adonis2 detects a system-wide difference, but the dataset is too small (5 sites per group) to localise it cleanly.
The picture sharpens with an ordination plot. NMDS projects the 30-dimensional species space into two axes that preserve rank distances, so you can literally see how groups separate.
The NMDS stress is 0.12, low enough to trust the 2-D projection. The ellipses show NM (Nature Management) sitting clearly to the left, BF and HF overlapping in the middle, and SF (Standard Farming) on the right. That visual matches the adonis2 result: Management explains a third of the spread, and NM is the most distinct group.
Try it: Build the same pairwise table using method = "BH" (Benjamini-Hochberg) instead of Bonferroni. Which pairs survive at FDR < 0.10?
Click to reveal solution
Explanation: Benjamini-Hochberg controls the false discovery rate rather than the family-wise error, so it has more power. At FDR < 0.10, two pairs involving NM survive, consistent with the NMDS picture.
Practice Exercises
These two capstones combine the concepts from above. Use distinct variable names so they do not overwrite the tutorial state.
Exercise 1: Two-factor PERMANOVA with R² breakdown
Fit a two-factor PERMANOVA on dune ~ Management + A1 (A1 is the soil thickness covariate) and report each term's R² and p-value as a tidy data frame named my_two_factor.
Click to reveal solution
Explanation: by = "terms" tests Management first, then A1 conditional on Management. The R² values add up to the explained variance: Management contributes 34%, A1 adds another 10% on top.
Exercise 2: Reusable PERMANOVA pipeline
Write a function my_permanova_pipeline(comm, group, dist = "bray") that takes a community matrix and a grouping factor, runs adonis2 and betadisper, and returns a one-row tibble with R2, permanova_p, and betadisper_p. Test it on dune and dune.env$Moisture.
Click to reveal solution
Explanation: The wrapper packages the workflow into a reusable function that returns just the numbers a paper or report needs. The pattern generalises to any community dataset.
Complete Example
Here is the full dune-Management pipeline collapsed into one runnable narrative. Each step is something you would do in a real ecology paper.
The five-step output is exactly what you would put in a methods paragraph: the omnibus PERMANOVA detects a Management effect (R² = 0.34, p = 0.003), dispersions are homogeneous (PERMDISP p = 0.15), and no individual pair survives Bonferroni correction at the n = 5-per-group sample size, suggesting the design needs more sites for cleanly localised contrasts.
Summary
The diagram below collapses the whole tutorial into one workflow card.
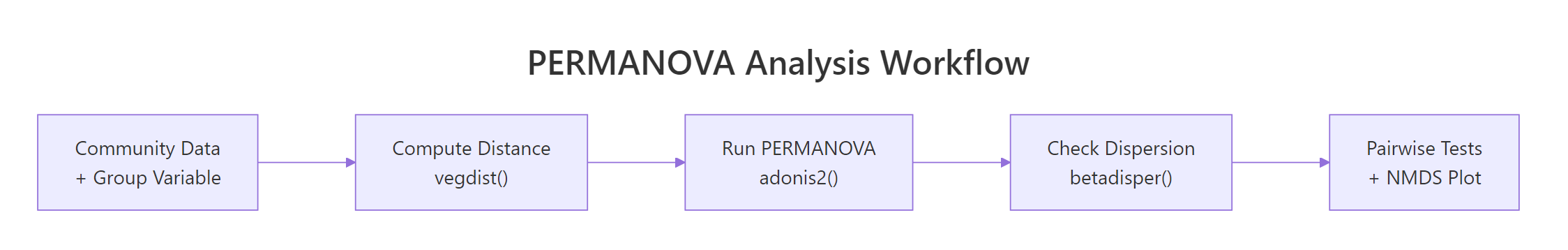
Figure 2: The end-to-end PERMANOVA workflow in vegan.
| Step | vegan function | What it answers |
|---|---|---|
| Compute distance | vegdist() |
How dissimilar are pairs of sites? |
| Test centroids | adonis2() |
Do groups differ overall? |
| Check spread | betadisper() + permutest() |
Is dispersion homogeneous across groups? |
| Localise differences | combn() loop + p.adjust() |
Which pairs of groups differ? |
| Visualise | metaMDS() + ordiplot() |
Does the picture match the test? |
Three things to take away. First, PERMANOVA is a permutation test on a distance matrix, not on raw values, so the metric you choose is part of the question. Second, the dispersion check is non-negotiable for unbalanced designs, run betadisper() before celebrating a small p-value. Third, always plot the ordination, the picture tells you whether a significant pseudo-F is biologically meaningful or just a centroid micro-shift.
References
- Anderson, M. J. (2001). A new method for non-parametric multivariate analysis of variance. Austral Ecology, 26(1), 32-46. Link
- McArdle, B. H., & Anderson, M. J. (2001). Fitting multivariate models to community data: a comment on distance-based redundancy analysis. Ecology, 82(1), 290-297. Link
- Oksanen, J. et al. vegan: Community Ecology Package (CRAN). Link
- vegan reference:
adonis2documentation. Link - Roberts, D. W. (2024). Applied Multivariate Statistics in R: PERMANOVA chapter. University of Washington. Link
- Anderson, M. J. (2014). Permutational Multivariate Analysis of Variance (PERMANOVA). Wiley StatsRef. Link
Continue Learning
- Bootstrap in R, the parent post on resampling-based inference, the same logic that powers permutation tests.
- Permutation Tests in R, the univariate permutation framework and natural prerequisite for PERMANOVA.
- When to Use Nonparametric Tests in R, a decision flowchart for picking a test when normality breaks down.