R Error: 'non-numeric argument to binary operator', Find the Hidden Character
Error in x * y : non-numeric argument to binary operator means R was asked to do arithmetic on something that isn't a number. The fastest way to find the culprit is to ask class() what each operand actually is, once you know that, the fix is usually a single conversion function.
What does "non-numeric argument to binary operator" mean in R?
R throws this error the moment an arithmetic operator, +, -, *, /, ^, %%, sees an operand that isn't numeric. The value may print exactly like a number and still be a character string, a factor, or a date under the hood. The one-line diagnosis is class(): once you know what R thinks each operand is, the fix is usually a single conversion function.
Let's reproduce the error on purpose, then solve it in three lines.
The string "19.99" and the number 19.99 print identically, but class() exposes the difference. Once you know which operand is the character, as.numeric() patches it in one call. The rest of this guide is about recognising the same mismatch when it's less obvious, buried inside data frames, hidden in whitespace, or wearing a factor disguise.
class() on every operand of the failing expression. The error almost always goes away once you know which side is not numeric.Try it: You have ex_price <- "42" and ex_qty <- 2. Write one line that computes the numeric total and stores it in ex_total.
Click to reveal solution
Explanation: as.numeric() converts the character "42" to the number 42, and R can then multiply it by ex_qty.
How do you find the guilty column inside a data frame?
When the error comes from a data frame expression like df$revenue * df$qty, the trick is to scan every column at once. R gives you three escalating tools: str() for a full dump, sapply(df, class) for a one-line class vector, and sapply(df, is.numeric) for a straight yes/no answer.
Let's build a tiny shop data frame with one obvious character column hiding among numeric ones, then find it three different ways.
str() is great for eyeballing, but sapply(shop, is.numeric) is what you actually want in a script, it returns a named logical vector you can feed straight into column selection. Wrapping it with names(shop)[!...] prints exactly the columns that would break an arithmetic expression.
names(df)[!sapply(df, is.numeric)]. Keep it in your snippet library, it's the single fastest way to find every non-numeric column in a data frame, no matter how wide.Try it: Using the data frame ex_df below, write one line that returns the names of every non-numeric column.
Click to reveal solution
Explanation: sapply(ex_df, is.numeric) returns TRUE for id and wins, FALSE for score and team. Negating it with ! flips the selection to the non-numeric columns.
What hidden characters silently turn a numeric-looking column into text?
The hardest version of this bug is the column that looks numeric when you print() it but fails is.numeric(). A single stray character anywhere in the column forces the whole thing to character. The usual suspects are:
- Leading or trailing whitespace,
" 19.99"from a poorly-trimmed CSV. - Currency symbols,
"$19.99"or"€12.00". - Thousand separators,
"1,200"from Excel exports. - Placeholder text,
"N/A","TBD","-". - Stray units,
"42kg","15%". - Unicode minus,
"−5"(U+2212) instead of ASCII-5.
The cleanest fix is readr::parse_number(), which strips everything that isn't part of a number and converts in one call. It also keeps the bad rows as NA instead of crashing, which is what you usually want.
parse_number() handled whitespace, $, ,, and "42kg" without a single manual gsub. The one value it couldn't rescue, "N/A", became NA, which sum(..., na.rm = TRUE) ignores. That's a much safer outcome than a hard error mid-script.
"N/A" or "$", the whole column becomes character and every arithmetic operation on it fails. Clean on import, not after the error hits.Try it: Clean the price vector ex_prices so its values become a numeric vector. Use any approach, gsub() + as.numeric() or parse_number().
Click to reveal solution
Explanation: parse_number() strips any non-numeric character automatically. The manual version uses gsub("\\$", "", ...) to remove the dollar signs, then as.numeric() to convert. Both produce the same numeric vector.
How do you fix the three common cause patterns?
Every instance of this error reduces to one of three root causes. Once you've diagnosed which pattern you're looking at, the fix is mechanical.
| Pattern | What it looks like | Fix |
|---|---|---|
| A. Character that looks numeric | "19.99" from CSV or user input |
as.numeric(x) |
| B. Factor used in arithmetic | factor(c(10, 20, 30)) |
as.numeric(as.character(x)) |
| C. Messy strings with noise | "$1,200", "42kg", " 19.99 " |
readr::parse_number(x) |
Let's apply all three to one small data frame so you can see the patterns side-by-side.
All three columns are numeric after the fix, and rowSums(mess), which would have errored before, now returns one total per row. Pattern B is the one that trips people up most: calling as.numeric() directly on a factor returns the level indices, not the label values. The as.character() step rescues the original strings first.
as.numeric(factor(c("10","20","30"))) returns 1 2 3, not 10 20 30. Always route factors through as.character() first: as.numeric(as.character(x)).Try it: ex_scores is a factor of test scores. Convert it to numeric and compute the mean.
Click to reveal solution
Explanation: as.character(ex_scores) recovers the label strings "88" "92" "79" "95" "84". as.numeric() then converts them to real numbers, and mean() averages them. Skipping as.character() would have averaged the level indices instead and produced the wrong answer.
Practice Exercises
Exercise 1: Clean an orders table and compute total cost
The orders data frame has two type problems: price is a character with dollar signs, and quantity is a factor. Compute total_cost = price * quantity for each row, then sum() them into a grand total called grand_total.
Click to reveal solution
Explanation: parse_number() strips the dollar signs from price in one call. quantity needs the as.character() detour because it's a factor. Once both columns are numeric, multiplication and sum() work.
Exercise 2: Auto-clean every non-numeric column in a survey
survey has three suspect columns (q1, q2, q3) that should all be numeric but look like they were typed by humans. Write a short pipeline that (a) finds the non-numeric columns, (b) cleans each with parse_number(), and (c) adds a row_total column. Don't hard-code column names, use sapply() so your code works on any survey.
Click to reveal solution
Explanation: sapply(survey, is.numeric) flags q1, q2, q3 as non-numeric, and we exclude respondent by name because it's supposed to stay character. lapply(..., parse_number) cleans them all in one pass. rowSums(..., na.rm = TRUE) tolerates the NA from the "N/A" cell so respondent B still gets a total.
Complete Example: Debug a Broken Revenue Report
Here's the kind of mess that shows up in real CSV exports. The revenue_df data frame should let us compute total revenue per region, but every numeric column is secretly a character, and one is a factor. Watch the diagnostic workflow end-to-end.
Four commands, four patches, one grand total. The key is that the sapply() scan in Step 1 told us exactly which columns needed which treatment before we wrote a single fix, no guessing, no re-running, no error messages to decode.
readr::read_csv(), pass col_types = cols(units_sold = col_number(), unit_price = col_number()) to parse numeric columns correctly at import time. Most of these errors never happen if the data enters R with the right types.Summary
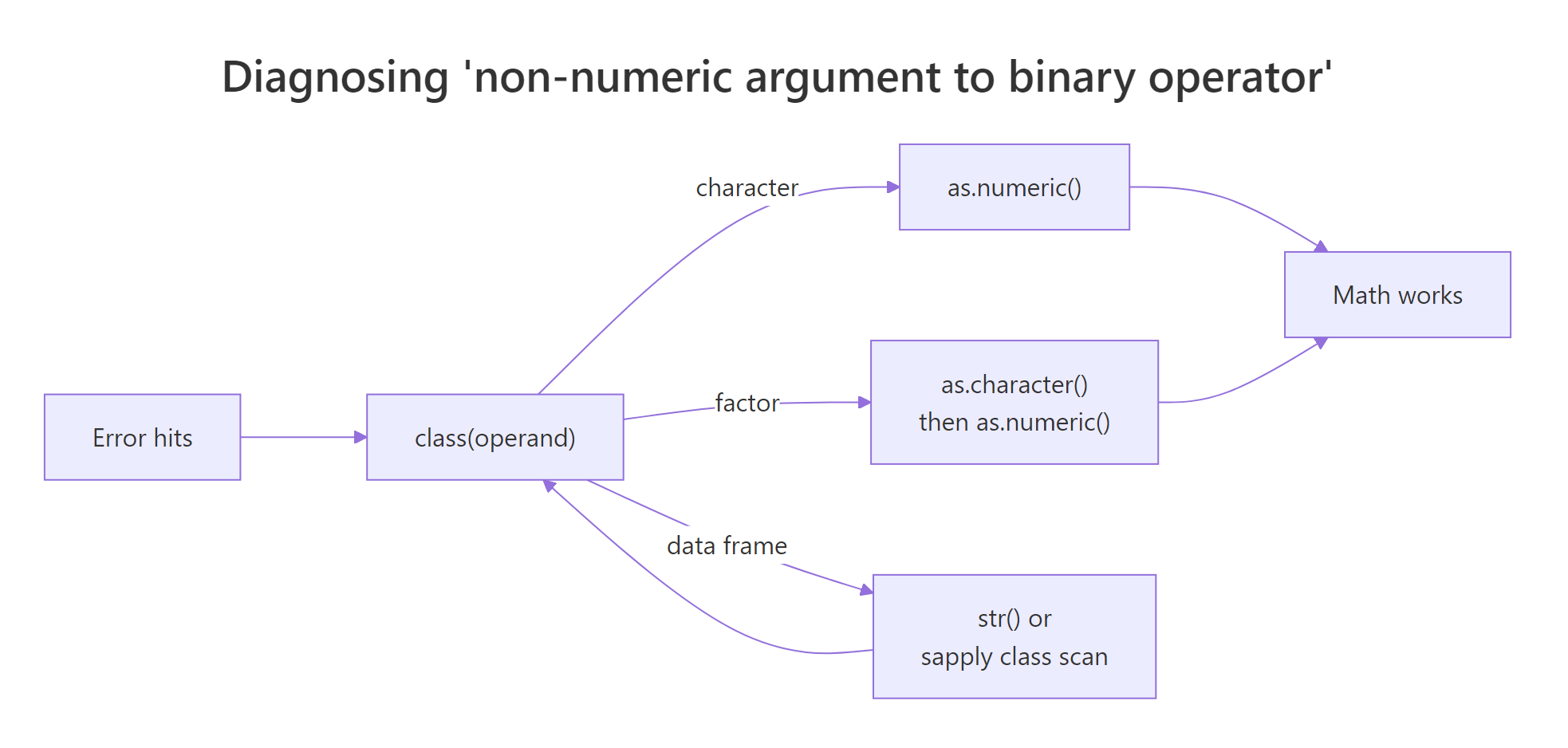
Figure 1: Diagnostic flow for the 'non-numeric argument to binary operator' error.
| Symptom | Diagnosis | Fix | Prevention |
|---|---|---|---|
Error on x * y |
class(x) returns "character" |
as.numeric(x) |
Check types after every data import |
Error on df$col * k |
sapply(df, is.numeric) flags the column |
parse_number() or as.numeric() |
Use col_types in read_csv() |
| Wrong numbers after converting a factor | class(x) returns "factor" |
as.numeric(as.character(x)) |
Avoid factors on numeric-looking columns |
Single NA after as.numeric() |
A hidden "N/A", "$", or whitespace |
parse_number(x, na = "N/A") |
Clean strings before arithmetic |
| Mystery failure in wide data | names(df)[!sapply(df, is.numeric)] |
Loop fixes with lapply() |
Validate with stopifnot(sapply(df, is.numeric)) |
References
- R Core Team, R Language Definition, Operators. Link
- Wickham, H., Advanced R (2nd ed.), Chapter 3: Vectors. Link
- readr,
parse_number()reference. Link - R Core Team, R FAQ:
as.numeric()on a factor. Link - R Documentation,
base::class(). Link - Wickham, H., Tidyverse Style Guide. Link
- Posit Community, "non-numeric argument to binary operator" thread. Link
Continue Learning
- R Errors Decoded: Plain-English Explanations and Exact Fixes, the parent reference for all 50 common R errors. R-Common-Errors.html
- R Warning: NAs introduced by coercion, what happens after
as.numeric()silently fails on bad values. R-Warning-NAs-Introduced-By-Coercion.html - R Error: object 'x' not found, when the variable name itself is the problem, not its type. R-Error-Object-Not-Found.html